The Antioxidant Cyclic 3-Hydroxymelatonin Promotes the Growth and Flowering of Arabidopsis thaliana
- PMID: 35740053
- PMCID: PMC9219689
- DOI: 10.3390/antiox11061157
The Antioxidant Cyclic 3-Hydroxymelatonin Promotes the Growth and Flowering of Arabidopsis thaliana
Abstract
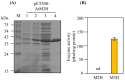
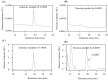
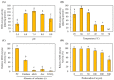
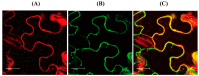
In plants, melatonin is metabolized into several compounds, including the potent antioxidant cyclic 3-hydroxymelatonin (3-OHM). Melatonin 3-hydroxylase (M3H), a member of the 2-oxo-glutarate-dependent enzyme family, is responsible for 3-OHM biosynthesis. Although rice M3H has been cloned, its roles are unclear, and no homologs in other plant species have been characterized. Here, we cloned and characterized Arabidopsis thaliana M3H (AtM3H). The purified recombinant AtM3H exhibited Km and Vmax values of 100 μM and 20.7 nmol/min/mg protein, respectively. M3H was localized to the cytoplasm, and its expression peaked at night. Based on a 1,1-diphenyl-2-picrylhydrazyl (DPPH) assay, 3-OHM exhibited 15-fold higher antioxidant activity than melatonin. An Arabidopsis M3H knockout mutant (m3h) produced less 3-OHM than the wildtype (WT), thus reducing antioxidant activity and biomass and delaying flowering. These defects were caused by reduced expression of FLOWERING LOCUS T (FT) and gibberellin-related genes, which are responsible for flowering and growth. Exogenous 3-OHM, but not exogenous melatonin, induced FT expression. The peak of M3H expression at night matched the FT expression pattern. The WT and m3h exhibited similar responses to salt stress and pathogens. Collectively, our findings indicate that 3-OHM promotes growth and flowering in Arabidopsis.
Keywords: Escherichia coli expression; FLOWERING LOCUS T; antioxidant; cyclic 3-hydroxymelatonin; gibberellin; melatonin; pathogen resistance; salt tolerance.
Conflict of interest statement
The authors declare no conflict of interest.
Figures






References
-
- Dobrikova A.G. Signaling molecules in plants: Exogenous application. Acta Sci. Agric. 2017;1:38–41.
-
- Lee H.Y., Back K. The phytomelatonin receptor (PMRT1) Arabidopsis Cand2 is not a bona fide G protein-coupled melatonin receptor. Melatonin Res. 2020;3:177–186. doi: 10.32794/mr11250055. - DOI
LinkOut - more resources
Full Text Sources
Research Materials

