Origin and Global Expansion of Mycobacterium tuberculosis Complex Lineage 3
- PMID: 35741753
- PMCID: PMC9222951
- DOI: 10.3390/genes13060990
Origin and Global Expansion of Mycobacterium tuberculosis Complex Lineage 3
Abstract
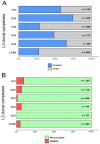
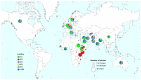
Mycobacterium tuberculosis complex (MTBC) Lineage 3 (L3) strains are abundant in world regions with the highest tuberculosis burden. To investigate the population structure and the global diversity of this major lineage, we analyzed a dataset comprising 2682 L3 strains from 38 countries over 5 continents, by employing 24-loci mycobacterial interspersed repetitive unit-variable number of tandem repeats genotyping (MIRU-VNTR) and drug susceptibility testing. We further combined whole-genome sequencing (WGS) and phylogeographic analysis for 373 strains representing the global L3 genetic diversity. Ancestral state reconstruction confirmed that the origin of L3 strains is located in Southern Asia and further revealed multiple independent introduction events into North-East and East Africa. This study provides a systematic understanding of the global diversity of L3 strains and reports phylogenetic variations that could inform clinical trials which evaluate the effectivity of new drugs/regimens or vaccine candidates.
Keywords: Lineage 3; MTBC; Mycobacterium tuberculosis; back to Africa.
Conflict of interest statement
P.S. is a consultant for Genoscreen. Other authors declare no conflict of interest.
Figures



References
-
- WHO . Global Tuberculosis Report. World Health Organization; Geneva, Switzerland: 2020. p. 232.
-
- Coscolla M., Gagneux S., Menardo F., Loiseau C., Ruiz-Rodriguez P., Borrell S., Otchere I.D., Asante-Poku A., Asare P., Sánchez-Busó L., et al. Phylogenomics of Mycobacterium africanum reveals a new lineage and a complex evolutionary history. Microb. Genom. 2021;7:000477. doi: 10.1099/mgen.0.000477. - DOI - PMC - PubMed

