NK Cells in Protection from HIV Infection
- PMID: 35746615
- PMCID: PMC9231282
- DOI: 10.3390/v14061143
NK Cells in Protection from HIV Infection
Abstract
Some people, known as HIV-exposed seronegative (HESN) individuals, remain uninfected despite high levels of exposure to HIV. Understanding the mechanisms underlying their apparent resistance to HIV infection may inform strategies designed to protect against HIV infection. Natural Killer (NK) cells are innate immune cells whose activation state depends on the integration of activating and inhibitory signals arising from cell surface receptors interacting with their ligands on neighboring cells. Inhibitory NK cell receptors use a subset of major histocompatibility (MHC) class I antigens as ligands. This interaction educates NK cells, priming them to respond to cells with reduced MHC class I antigen expression levels as occurs on HIV-infected cells. NK cells can interact with both autologous HIV-infected cells and allogeneic cells bearing MHC antigens seen as non self by educated NK cells. NK cells are rapidly activated upon interacting with HIV-infected or allogenic cells to elicit anti-viral activity that blocks HIV spread to new target cells, suppresses HIV replication, and kills HIV-infected cells before HIV reservoirs can be seeded and infection can be established. In this manuscript, we will review the epidemiological and functional evidence for a role for NK cells in protection from HIV infection.
Keywords: HIV; HIV-exposed seronegative; HLA; HLA-B*57; killer immunoglobulin-like receptors; natural killer cells.
Conflict of interest statement
The authors declare no conflict of interest.
Figures

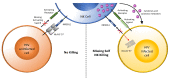
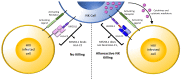
Similar articles
-
The Education of NK Cells Determines Their Responsiveness to Autologous HIV-Infected CD4 T Cells.J Virol. 2019 Nov 13;93(23):e01185-19. doi: 10.1128/JVI.01185-19. Print 2019 Dec 1. J Virol. 2019. PMID: 31511383 Free PMC article.
-
Natural Killer Cells in Antibody Independent and Antibody Dependent HIV Control.Front Immunol. 2022 May 20;13:879124. doi: 10.3389/fimmu.2022.879124. eCollection 2022. Front Immunol. 2022. PMID: 35720328 Free PMC article. Review.
-
Expression Profiles of Ligands for Activating Natural Killer Cell Receptors on HIV Infected and Uninfected CD4⁺ T Cells.Viruses. 2017 Oct 12;9(10):295. doi: 10.3390/v9100295. Viruses. 2017. PMID: 29023371 Free PMC article.
-
Natural killer (NK) cell receptor-HLA ligand genotype combinations associated with protection from HIV infection: investigation of how protective genotypes influence anti HIV NK cell functions.AIDS Res Ther. 2017 Sep 12;14(1):38. doi: 10.1186/s12981-017-0172-9. AIDS Res Ther. 2017. PMID: 28893287 Free PMC article. Review.
-
HLA-F on Autologous HIV-Infected Cells Activates Primary NK Cells Expressing the Activating Killer Immunoglobulin-Like Receptor KIR3DS1.J Virol. 2019 Aug 28;93(18):e00933-19. doi: 10.1128/JVI.00933-19. Print 2019 Sep 15. J Virol. 2019. PMID: 31270222 Free PMC article.
Cited by
-
Innate Immune Cell Functions Contribute to Spontaneous HIV Control.Curr HIV/AIDS Rep. 2024 Nov 30;22(1):6. doi: 10.1007/s11904-024-00713-0. Curr HIV/AIDS Rep. 2024. PMID: 39614998 Free PMC article. Review.
-
Recombinant Human Interleukin-2 Corrects NK Cell Phenotype and Functional Activity in Patients with Post-COVID Syndrome.Pharmaceuticals (Basel). 2023 Apr 3;16(4):537. doi: 10.3390/ph16040537. Pharmaceuticals (Basel). 2023. PMID: 37111294 Free PMC article.
-
Fragment-Based Immune Cell Engager Antibodies in Treatment of Cancer, Infectious and Autoimmune Diseases: Lessons and Insights from Clinical and Translational Studies.Antibodies (Basel). 2025 Jun 24;14(3):52. doi: 10.3390/antib14030052. Antibodies (Basel). 2025. PMID: 40700292 Free PMC article. Review.
-
Analysis of KIR and HLA Polymorphism in Chinese Individuals With COVID-19.HLA. 2024 Oct;104(4):e15715. doi: 10.1111/tan.15715. HLA. 2024. PMID: 39364548
References
-
- Beretta A., Furci L., Burastero S., Cosma A., Dinelli M.E., Lopalco L., DeSantis C., Tambussi G., Carrow E., Sabbatani S., et al. HIV-1-specific immunity in persistently seronegative individuals at high risk for HIV infection. Immunol. Lett. 1996;51:39–43. doi: 10.1016/0165-2478(96)02553-9. - DOI - PubMed
-
- Clerici M., Giorgi J.V., Chou C.C., Gudeman V.K., Zack J.A., Gupta P., Ho H.N., Nishanian P.G., Berzofsky J.A., Shearer G.M. Cell-mediated immune response to human immunodeficiency virus (HIV) type 1 in seronegative homosexual men with recent sexual exposure to HIV-1. J. Infect. Dis. 1992;165:1012–1019. doi: 10.1093/infdis/165.6.1012. - DOI - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Medical
Research Materials

