Functional Analysis of a Frontal miRNA Cluster Located in the Large Latency Transcript of Pseudorabies Virus
- PMID: 35746619
- PMCID: PMC9227234
- DOI: 10.3390/v14061147
Functional Analysis of a Frontal miRNA Cluster Located in the Large Latency Transcript of Pseudorabies Virus
Abstract
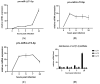
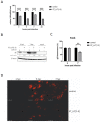
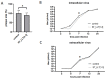
MicroRNAs (miRNAs) have been identified as a class of crucial regulators of virus-host crosstalk, modulating such processes as viral replication, antiviral immune response, viral latency, and pathogenesis. Pseudorabies virus (PRV), a model for the study of alphaherpesvirus biology, codes for 11 distinct miRNAs mapped to the ~4.6 kb intron of Large Latency Transcript (LLT). Recent studies have revealed the role of clusters consisting of nine and eleven miRNA genes in the replication and virulence of PRV. The function of separate miRNA species in regulating PRV biology has not been thoroughly investigated. To analyze the regulatory potential of three PRV miRNAs located in the frontal cluster of the LLT intron, we generated a research model based on the constitutive expression of viral miRNAs in swine testis cells (ST_LLT [1-3] cell line). Using a cell culture system providing a stable production of individual miRNAs at high levels, we demonstrated that the LLT [1-3] miRNA cluster significantly downregulated IE180, EP0, and gE at the early stages of PRV infection. It was further determined that LLT [1-3] miRNAs could regulate the infection process, leading to a slight distortion in transmission and proliferation ability. Collectively, our findings indicate the potential of LLT [1-3] miRNAs to retard the host responses by reducing viral antigenic load and suppressing the expansion of progeny viruses at the early stages of infection.
Keywords: EP0; IE180; Large Latency Transcript; glycoprotein E; miRNA cluster; pseudorabies virus.
Conflict of interest statement
The authors have no conflict of interest to disclose.
Figures


Similar articles
-
The full-length microRNA cluster in the intron of large latency transcript is associated with the virulence of pseudorabies virus.Virology. 2018 Jul;520:59-66. doi: 10.1016/j.virol.2018.05.004. Epub 2018 May 16. Virology. 2018. PMID: 29777914
-
Pseudorabies virus infected porcine epithelial cell line generates a diverse set of host microRNAs and a special cluster of viral microRNAs.PLoS One. 2012;7(1):e30988. doi: 10.1371/journal.pone.0030988. Epub 2012 Jan 23. PLoS One. 2012. PMID: 22292087 Free PMC article.
-
A 2.5-kilobase deletion containing a cluster of nine microRNAs in the latency-associated-transcript locus of the pseudorabies virus affects the host response of porcine trigeminal ganglia during established latency.J Virol. 2015 Jan;89(1):428-42. doi: 10.1128/JVI.02181-14. Epub 2014 Oct 15. J Virol. 2015. PMID: 25320324 Free PMC article.
-
A Comparison of Pseudorabies Virus Latency to Other α-Herpesvirinae Subfamily Members.Viruses. 2022 Jun 24;14(7):1386. doi: 10.3390/v14071386. Viruses. 2022. PMID: 35891367 Free PMC article. Review.
-
Molecular biology of pseudorabies virus: impact on neurovirology and veterinary medicine.Microbiol Mol Biol Rev. 2005 Sep;69(3):462-500. doi: 10.1128/MMBR.69.3.462-500.2005. Microbiol Mol Biol Rev. 2005. PMID: 16148307 Free PMC article. Review.
Cited by
-
Proteomic profiling of pseudorabies virus-infected PK-15 cells based on 4D label free analysis.Vet Res Forum. 2025;16(3):141-147. doi: 10.30466/vrf.2024.2026963.4239. Epub 2025 Mar 15. Vet Res Forum. 2025. PMID: 40391135 Free PMC article.
-
Global trends in research on miRNA-microbiome interaction from 2011 to 2021: A bibliometric analysis.Front Pharmacol. 2022 Aug 30;13:974741. doi: 10.3389/fphar.2022.974741. eCollection 2022. Front Pharmacol. 2022. PMID: 36110534 Free PMC article.
References
-
- Mettenleiter T.C. Pseudorabies (Aujeszky’s disease) virus: State of the art. August 1993. Acta Vet. Hung. 1994;42:153–177. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources

