Predicting RNA distance-based contact maps by integrated deep learning on physics-inferred secondary structure and evolutionary-derived mutational coupling
- PMID: 35751593
- PMCID: PMC9364379
- DOI: 10.1093/bioinformatics/btac421
Predicting RNA distance-based contact maps by integrated deep learning on physics-inferred secondary structure and evolutionary-derived mutational coupling
Abstract
Motivation: Recently, AlphaFold2 achieved high experimental accuracy for the majority of proteins in Critical Assessment of Structure Prediction (CASP 14). This raises the hope that one day, we may achieve the same feat for RNA structure prediction for those structured RNAs, which is as fundamentally and practically important similar to protein structure prediction. One major factor in the recent advancement of protein structure prediction is the highly accurate prediction of distance-based contact maps of proteins.
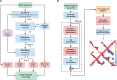
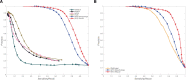
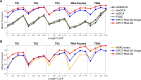
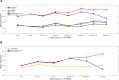
Results: Here, we showed that by integrated deep learning with physics-inferred secondary structures, co-evolutionary information and multiple sequence-alignment sampling, we can achieve RNA contact-map prediction at a level of accuracy similar to that in protein contact-map prediction. More importantly, highly accurate prediction for top L long-range contacts can be assured for those RNAs with a high effective number of homologous sequences (Neff > 50). The initial use of the predicted contact map as distance-based restraints confirmed its usefulness in 3D structure prediction.
Availability and implementation: SPOT-RNA-2D is available as a web server at https://sparks-lab.org/server/spot-rna-2d/ and as a standalone program at https://github.com/jaswindersingh2/SPOT-RNA-2D.
Supplementary information: Supplementary data are available at Bioinformatics online.
© The Author(s) 2022. Published by Oxford University Press.
Figures

References
-
- Ba J.L. et al. (2016) Layer normalization. Preprint arXiv, 1607.06450.
-
- Balakrishnan S. et al. (2011) Learning generative models for protein fold families. Proteins Struct. Funct. Bioinform., 79, 1061–1078. - PubMed
-
- Cai Z. et al. (2020) RIC-seq for global in situ profiling of RNA–RNA spatial interactions. Nature, 582, 432–437. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Miscellaneous

