The landscape of submicroscopic structural variants at the OPN1LW/OPN1MW gene cluster on Xq28 underlying blue cone monochromacy
- PMID: 35759666
- PMCID: PMC9271157
- DOI: 10.1073/pnas.2115538119
The landscape of submicroscopic structural variants at the OPN1LW/OPN1MW gene cluster on Xq28 underlying blue cone monochromacy
Abstract
Blue cone monochromacy (BCM) is an X-linked retinal disorder characterized by low vision, photoaversion, and poor color discrimination. BCM is due to the lack of long-wavelength-sensitive and middle-wavelength-sensitive cone photoreceptor function and caused by mutations in the OPN1LW/OPN1MW gene cluster on Xq28. Here, we investigated the prevalence and the landscape of submicroscopic structural variants (SVs) at single-base resolution in BCM patients. We found that about one-third (n = 73) of the 213 molecularly confirmed BCM families carry an SV, most commonly deletions restricted to the OPN1LW/OPN1MW gene cluster. The structure and precise breakpoints of the SVs were resolved in all but one of the 73 families. Twenty-two families-all from the United States-showed the same SV, and we confirmed a common ancestry of this mutation. In total, 42 distinct SVs were identified, including 40 previously unreported SVs, thereby quadrupling the number of precisely mapped SVs underlying BCM. Notably, there was no "region of overlap" among these SVs. However, 90% of SVs encompass the upstream locus control region, an essential enhancer element. Its minimal functional extent based on deletion mapping in patients was refined to 358 bp. Breakpoint analyses suggest diverse mechanisms underlying SV formation as well as in one case the gene conversion-based exchange of a 142-bp deletion between opsin genes. Using parsimonious assumptions, we reconstructed the composition and copy number of the OPN1LW/OPN1MW gene cluster prior to the mutation event and found evidence that large gene arrays may be predisposed to the occurrence of SVs at this locus.
Keywords: BCM; gene conversion; human visual pigment genes; locus control region; opsin gene deletion.
Conflict of interest statement
The authors declare no competing interest.
Figures

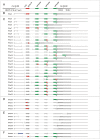

References
-
- Nathans J., et al. , Molecular genetics of human blue cone monochromacy. Science 245, 831–838 (1989). - PubMed
-
- Nathans J., Thomas D., Hogness D. S., Molecular genetics of human color vision: The genes encoding blue, green, and red pigments. Science 232, 193–202 (1986a). - PubMed
-
- Nathans J., Piantanida T. P., Eddy R. L., Shows T. B., Hogness D. S., Molecular genetics of inherited variation in human color vision. Science 232, 203–210 (1986b). - PubMed
Publication types
MeSH terms
Substances
Supplementary concepts
Grants and funding
LinkOut - more resources
Full Text Sources
Medical
Molecular Biology Databases
Research Materials

