Mechanisms orchestrating the enzymatic activity and cellular functions of deubiquitinases
- PMID: 35764170
- PMCID: PMC9356280
- DOI: 10.1016/j.jbc.2022.102198
Mechanisms orchestrating the enzymatic activity and cellular functions of deubiquitinases
Abstract
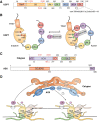
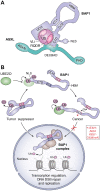
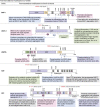
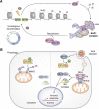
Deubiquitinases (DUBs) are required for the reverse reaction of ubiquitination and act as major regulators of ubiquitin signaling processes. Emerging evidence suggests that these enzymes are regulated at multiple levels in order to ensure proper and timely substrate targeting and to prevent the adverse consequences of promiscuous deubiquitination. The importance of DUB regulation is highlighted by disease-associated mutations that inhibit or activate DUBs, deregulating their ability to coordinate cellular processes. Here, we describe the diverse mechanisms governing protein stability, enzymatic activity, and function of DUBs. In particular, we outline how DUBs are regulated by their protein domains and interacting partners. Intramolecular interactions can promote protein stability of DUBs, influence their subcellular localization, and/or modulate their enzymatic activity. Remarkably, these intramolecular interactions can induce self-deubiquitination to counteract DUB ubiquitination by cognate E3 ubiquitin ligases. In addition to intramolecular interactions, DUBs can also oligomerize and interact with a wide variety of cellular proteins, thereby forming obligate or facultative complexes that regulate their enzymatic activity and function. The importance of signaling and post-translational modifications in the integrated control of DUB function will also be discussed. While several DUBs are described with respect to the multiple layers of their regulation, the tumor suppressor BAP1 will be outlined as a model enzyme whose localization, stability, enzymatic activity, and substrate recognition are highly orchestrated by interacting partners and post-translational modifications.
Keywords: BAP1; DUB activity; deubiquitinase; folding; multiprotein complex; post-translational modifications; quality control; ubiquitin.
Crown Copyright © 2022. Published by Elsevier Inc. All rights reserved.
Conflict of interest statement
Conflicts of interest The authors declare that they have no conflicts of interest with the content of this article.
Figures




Similar articles
-
Friend or foe? Reciprocal regulation between E3 ubiquitin ligases and deubiquitinases.Biochem Soc Trans. 2024 Feb 28;52(1):241-267. doi: 10.1042/BST20230454. Biochem Soc Trans. 2024. PMID: 38414432 Free PMC article. Review.
-
Deubiquitinases in cancer.Nat Rev Cancer. 2023 Dec;23(12):842-862. doi: 10.1038/s41568-023-00633-y. Epub 2023 Nov 7. Nat Rev Cancer. 2023. PMID: 37935888 Review.
-
The emerging roles of deubiquitinases in plant proteostasis.Essays Biochem. 2022 Aug 5;66(2):147-154. doi: 10.1042/EBC20210060. Essays Biochem. 2022. PMID: 35678302 Free PMC article. Review.
-
Deubiquitinases: Modulators of Different Types of Regulated Cell Death.Int J Mol Sci. 2021 Apr 21;22(9):4352. doi: 10.3390/ijms22094352. Int J Mol Sci. 2021. PMID: 33919439 Free PMC article. Review.
-
Deubiquitinating enzymes (DUBs): Regulation, homeostasis, and oxidative stress response.J Biol Chem. 2021 Sep;297(3):101077. doi: 10.1016/j.jbc.2021.101077. Epub 2021 Aug 12. J Biol Chem. 2021. PMID: 34391779 Free PMC article. Review.
Cited by
-
Targeting the Ubiquitin-Proteasome System and Recent Advances in Cancer Therapy.Cells. 2023 Dec 22;13(1):29. doi: 10.3390/cells13010029. Cells. 2023. PMID: 38201233 Free PMC article. Review.
-
Therapeutic Targeting of the GLS1-c-Myc Positive Feedback Loop Suppresses Glutaminolysis and Inhibits Progression of Head and Neck Cancer.Cancer Res. 2024 Oct 1;84(19):3223-3234. doi: 10.1158/0008-5472.CAN-24-0254. Cancer Res. 2024. PMID: 39024547 Free PMC article.
-
TNF receptor-associated factors: promising targets of natural products for the treatment of osteoporosis.Front Physiol. 2025 May 27;16:1527814. doi: 10.3389/fphys.2025.1527814. eCollection 2025. Front Physiol. 2025. PMID: 40496246 Free PMC article. Review.
-
Elucidating the role of ubiquitination and deubiquitination in osteoarthritis progression.Front Immunol. 2023 Jun 9;14:1217466. doi: 10.3389/fimmu.2023.1217466. eCollection 2023. Front Immunol. 2023. PMID: 37359559 Free PMC article. Review.
-
Deaggregation of mutant Plasmodium yoelii de-ubiquitinase UBP1 alters MDR1 localization to confer multidrug resistance.Nat Commun. 2024 Feb 27;15(1):1774. doi: 10.1038/s41467-024-46006-3. Nat Commun. 2024. PMID: 38413566 Free PMC article.
References
-
- Komander D., Rape M. The ubiquitin code. Annu. Rev. Biochem. 2012;81:203–229. - PubMed
-
- Yau R., Rape M. The increasing complexity of the ubiquitin code. Nat. Cell Biol. 2016;18:579–586. - PubMed
-
- Rape M. Ubiquitylation at the crossroads of development and disease. Nat. Rev. Mol. Cell Biol. 2018;19:59–70. - PubMed
-
- Popovic D., Vucic D., Dikic I. Ubiquitination in disease pathogenesis and treatment. Nat. Med. 2014;20:1242–1253. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

