Multiplexed Ultrasound Imaging Using Spectral Analysis on Gas Vesicles
- PMID: 35765741
- PMCID: PMC9463101
- DOI: 10.1002/adhm.202200568
Multiplexed Ultrasound Imaging Using Spectral Analysis on Gas Vesicles
Abstract
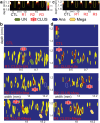
Current advances in ultrasound imaging techniques combined with the next generation contrast agents such as gas vesicles (GV) revolutionize the visualization of biological tissues with spatiotemporal precision. In optics, fluorescent proteins enable understanding of molecular and cellular functions in biological systems due to their multiplexed imaging capability. Here, a panel of GVs is investigated using mid-band fit (MBF) spectral imaging to realize multiplexed ultrasound imaging to uniquely visualize locations of different types of stationary GVs. The MBF spectral imaging technique demonstrates that stationary clustered GVs are efficiently localized and distinguished from unclustered GVs in agarose gel phantom and 3D vessel structures are visualized in ex vivo mouse liver specimens. Mouse macrophages serve as carriers of clustered and unclustered GVs and multiplexing beacons to report cells' spatial locations by emitting distinct spectral signals. 2D MBF spectral images are reconstructed, and pixels in these images are classified depending on MBF values by comparing predetermined filters that predict the existence of cells with clustered and unclustered GVs. This pseudo-coloring scheme clearly distinguishes the locations of two classes of cells like pseudo-color images in fluorescence microscopy.
Keywords: Serratia sp. ATCC 39006; clustered gas vesicles; high frequency ultrasound; mid-band fit spectral imaging; multiplexed ultrasound imaging; pseudo-coloring; ultrasound contrast agents.
© 2022 The Authors. Advanced Healthcare Materials published by Wiley-VCH GmbH.
Conflict of interest statement
The authors declare no conflict of interest.
Figures






Similar articles
-
Genetic Modulation of Biosynthetic Gas Vesicles for Ultrasound Imaging.Small. 2024 Aug;20(33):e2310008. doi: 10.1002/smll.202310008. Epub 2024 Mar 27. Small. 2024. PMID: 38533968
-
Nonlinear X-wave ultrasound imaging of acoustic biomolecules.Phys Rev X. 2018 Oct-Dec;8(4):041002. doi: 10.1103/physrevx.8.041002. Epub 2018 Oct 4. Phys Rev X. 2018. PMID: 34040818 Free PMC article.
-
Geometric effects in gas vesicle buckling under ultrasound.Biophys J. 2022 Nov 1;121(21):4221-4228. doi: 10.1016/j.bpj.2022.09.004. Epub 2022 Sep 7. Biophys J. 2022. PMID: 36081347 Free PMC article.
-
The latest in ultrasound: three-dimensional imaging. Part II.Eur J Radiol. 1998 May;27 Suppl 2:S183-7. doi: 10.1016/s0720-048x(98)00077-1. Eur J Radiol. 1998. PMID: 9652520 Review.
-
Advances in the application of gas vesicles in medical imaging and disease treatment.J Biol Eng. 2024 Jul 23;18(1):41. doi: 10.1186/s13036-024-00426-3. J Biol Eng. 2024. PMID: 39044273 Free PMC article. Review.
Cited by
-
Gas Vesicle-Blood Interactions Enhance Ultrasound Imaging Contrast.Nano Lett. 2023 Dec 13;23(23):10748-10757. doi: 10.1021/acs.nanolett.3c02780. Epub 2023 Nov 20. Nano Lett. 2023. PMID: 37983479 Free PMC article.
-
Ultrasound-Mediated Membrane Modulation for Biomedical Applications.Nanomaterials (Basel). 2025 Jun 7;15(12):884. doi: 10.3390/nano15120884. Nanomaterials (Basel). 2025. PMID: 40559247 Free PMC article. Review.
References
-
- a) Lizzi F. L., Astor M., Feleppa E. J., Shao M., Kalisz A., Ultrasound Med. Biol. 1997, 23, 1371; - PubMed
- b) Han A., Abuhabsah R., Blue J. P. Jr., Sarwate S., O'Brien W. D. Jr., J. Acoust. Soc. Am. 2011, 130, 4139; - PMC - PubMed
- c) Xu G., Fowlkes J. B., Tao C., Liu X., Wang X., Ultrasound Med. Biol. 2015, 41, 1473. - PMC - PubMed
-
- a) Fuentes V. L., Moran C. M., Schober K., McEwan J. D., Brown H., Sutherland G. R., McDicken W. N., Am. J. Vet. Res. 1997, 58, 1055; - PubMed
- b) Pawlicki A. D., O'Brien W. D. Jr., Ultrason. Imaging 2013, 35, 162; - PMC - PubMed
- c) Boozari B., Potthoff A., Mederacke I., Hahn A., Reising A., Rifai K., Wedemeyer H., Bahr M., Kubicka S., Manns M., Gebel M., J. Ultrasound Med. 2010, 29, 1581; - PubMed
- d) Padilla F., Jenson F., Laugier P., Ultrasonics 2006, 44, e57; - PubMed
- e) Lizzi F. L., Ostromogilsky M., Feleppa E. J., Rorke M. C., Yaremko M. M., IEEE Trans. Ultrason. Ferroelectr. Freq. Control 1987, 34, 319. - PubMed
-
- Lizzi F. L., Greenebaum M., Feleppa E. J., Elbaum M., Coleman D. J., J. Acoust. Soc. Am. 1983, 73, 1366. - PubMed
-
- Chalfie M., Tu Y., Euskirchen G., Ward W. W., Prasher D. C., Science 1994, 263, 802. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

