Core transcription programs controlling injury-induced neurodegeneration of retinal ganglion cells
- PMID: 35767995
- PMCID: PMC9391318
- DOI: 10.1016/j.neuron.2022.06.003
Core transcription programs controlling injury-induced neurodegeneration of retinal ganglion cells
Erratum in
-
Core transcription programs controlling injury-induced neurodegeneration of retinal ganglion cells.Neuron. 2023 Feb 1;111(3):444. doi: 10.1016/j.neuron.2023.01.008. Neuron. 2023. PMID: 36731430 Free PMC article. No abstract available.
Abstract
Regulatory programs governing neuronal death and axon regeneration in neurodegenerative diseases remain poorly understood. In adult mice, optic nerve crush (ONC) injury by severing retinal ganglion cell (RGC) axons results in massive RGC death and regenerative failure. We performed an in vivo CRISPR-Cas9-based genome-wide screen of 1,893 transcription factors (TFs) to seek repressors of RGC survival and axon regeneration following ONC. In parallel, we profiled the epigenetic and transcriptional landscapes of injured RGCs by ATAC-seq and RNA-seq to identify injury-responsive TFs and their targets. These analyses converged on four TFs as critical survival regulators, of which ATF3/CHOP preferentially regulate pathways activated by cytokines and innate immunity and ATF4/C/EBPγ regulate pathways engaged by intrinsic neuronal stressors. Manipulation of these TFs protects RGCs in a glaucoma model. Our results reveal core transcription programs that transform an initial axonal insult into a degenerative process and suggest novel strategies for treating neurodegenerative diseases.
Keywords: ATF3; ATF4; C/EBPg; CHOP; axon regeneration and neuronal degeneration; optic nerve; retinal ganglion cell.
Copyright © 2022 Elsevier Inc. All rights reserved.
Conflict of interest statement
Declaration of interests C.J.W. is a founder of Nocion Therapeutics and QurAlis. J.R.S. is a consultant for Biogen. Z.H. is an advisor of SpineX, Life Biosciences, and Myro Therapeutics.
Figures





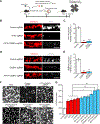

Comment in
-
Live, die, or regenerate? New insights from multi-omic analyses.Neuron. 2022 Aug 17;110(16):2516-2519. doi: 10.1016/j.neuron.2022.07.026. Neuron. 2022. PMID: 35981522 Free PMC article.
References
-
- Abel T, Martin KC, Bartsch D, and Kandel ER (1998). Memory suppressor genes: inhibitory constraints on the storage of long-term memory. Science 279, 338–341. - PubMed
-
- Aguayo AJ, Rasminsky M, Bray GM, Carbonetto S, McKerracher L, Villegas-Perez MP, Vidal-Sanz M, and Carter DA (1991). Degenerative and regenerative responses of injured neurons in the central nervous system of adult mammals. Philos Trans R Soc Lond B Biol Sci 331, 337–343. - PubMed
-
- Almasieh M, and Levin LA (2017). Neuroprotection in Glaucoma: Animal Models and Clinical Trials. Annu Rev Vis Sci 3, 91–120. - PubMed
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Medical
Molecular Biology Databases
Research Materials
Miscellaneous

