Distinct gene expression dynamics in developing and regenerating crustacean limbs
- PMID: 35776546
- PMCID: PMC9271199
- DOI: 10.1073/pnas.2119297119
Distinct gene expression dynamics in developing and regenerating crustacean limbs
Abstract
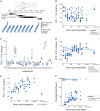
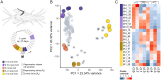
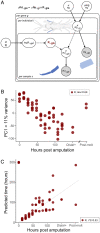
Regenerating animals have the ability to reproduce body parts that were originally made in the embryo and subsequently lost due to injury. Understanding whether regeneration mirrors development is an open question in most regenerative species. Here, we take a transcriptomics approach to examine whether leg regeneration shows similar temporal patterns of gene expression as leg development in the embryo, in the crustacean Parhyale hawaiensis. We find that leg development in the embryo shows stereotypic temporal patterns of gene expression. In contrast, the dynamics of gene expression during leg regeneration show a higher degree of variation related to the physiology of individual animals. A major driver of this variation is the molting cycle. We dissect the transcriptional signals of individual physiology and regeneration to obtain clearer temporal signals marking distinct phases of leg regeneration. Comparing the transcriptional dynamics of development and regeneration we find that, although the two processes use similar sets of genes, the temporal patterns in which these genes are deployed are different and cannot be systematically aligned.
Keywords: Parhyale hawaiensis; crustacean; leg development; regeneration; transcriptional profiling.
Conflict of interest statement
The authors declare no competing interest.
Figures



References
-
- Skinner D. M., “Molting and regeneration.” in The Biology of Crustacea: Integument, Pigments, and Hormonal Processes, Bliss D. E., Mantel L. H., Eds. (Academic Press, 1985) pp. 43–146.
-
- Singer M., The influence of the nerve in regeneration of the amphibian extremity. Q. Rev. Biol. 27, 169–200 (1952). - PubMed
-
- Kyritsis N., et al. , Acute inflammation initiates the regenerative response in the adult zebrafish brain. Science 338, 1353–1356 (2012). - PubMed
Publication types
MeSH terms
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

