Molecular mechanism of S-adenosylmethionine sensing by SAMTOR in mTORC1 signaling
- PMID: 35776786
- PMCID: PMC10883374
- DOI: 10.1126/sciadv.abn3868
Molecular mechanism of S-adenosylmethionine sensing by SAMTOR in mTORC1 signaling
Abstract
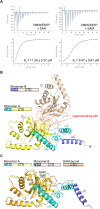
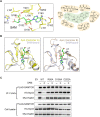
The mechanistic target of rapamycin-mLST8-raptor complex (mTORC1) functions as a central regulator of cell growth and metabolism in response to changes in nutrient signals such as amino acids. SAMTOR is an S-adenosylmethionine (SAM) sensor, which regulates the mTORC1 activity through its interaction with the GTPase-activating protein activity toward Rags-1 (GATOR1)-KPTN, ITFG2, C12orf66 and SZT2-containing regulator (KICSTOR) complex. In this work, we report the crystal structures of Drosophila melanogaster SAMTOR in apo form and in complex with SAM. SAMTOR comprises an N-terminal helical domain and a C-terminal SAM-dependent methyltransferase (MTase) domain. The MTase domain contains the SAM-binding site and the potential GATOR1-KICSTOR-binding site. The helical domain functions as a molecular switch, which undergoes conformational change upon SAM binding and thereby modulates the interaction of SAMTOR with GATOR1-KICSTOR. The functional roles of the key residues and the helical domain are validated by functional assays. Our structural and functional data together reveal the molecular mechanism of the SAM sensing of SAMTOR and its functional role in mTORC1 signaling.
Figures


References
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

