Collagen VII maintains proteostasis in dermal fibroblasts by scaffolding TANGO1 cargo
- PMID: 35779741
- PMCID: PMC9683098
- DOI: 10.1016/j.matbio.2022.06.008
Collagen VII maintains proteostasis in dermal fibroblasts by scaffolding TANGO1 cargo
Abstract
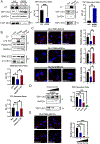
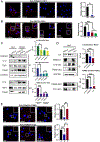
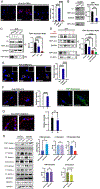
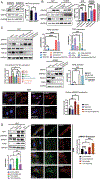
Lack of type VII collagen (C7) disrupts cellular proteostasis yet the mechanism remains undescribed. By studying the relationship between C7 and the extracellular matrix (ECM)-associated proteins thrombospondin-1 (TSP1), type XII collagen (C12) and tissue transglutaminase (TGM2) in primary human dermal fibroblasts from multiple donors with or without the genetic disease recessive dystrophic epidermolysis bullosa (RDEB) (n=31), we demonstrate that secretion of each of these proteins is increased in the presence of C7. In dermal fibroblasts isolated from patients with RDEB, where C7 is absent or defective, association with the COPII outer coat protein SEC31 and ultimately secretion of each of these ECM-associated proteins is reduced and intracellular levels are increased. In RDEB fibroblasts, overall collagen secretion (as determined by the levels of hydroxyproline in the media) is unchanged while traffic from the ER to Golgi of TSP1, C12 and TGM2 occurs in a type I collagen (C1) dependent manner. In normal fibroblasts association of TSP1, C12 and TGM2 with the ER exit site transmembrane protein Transport ANd Golgi Organization-1 (TANGO1) as determined by proximity ligation assays, requires C7. In the absence of wild-type C7, or when ECM-associated proteins are overexpressed, C1 proximity and intracellular levels increase resulting in elevated cellular stress responses and elevated TGFβ signaling. Collectively, these data demonstrate a role for C7 in loading COPII vesicle cargo and provides a mechanism for disrupted proteostasis, elevated cellular stress and increased TGFβ signaling in patients with RDEB. Furthermore, our data point to a threshold of cargo loading that can be exceeded with increased protein levels leading to pathological outcomes in otherwise normal cells.
Keywords: Collagen VII; ER stress; TANGO1; and TGFβ signaling; recessive dystrophic epidermolysis bullosa; thrombospondin.
Copyright © 2022 The Authors. Published by Elsevier B.V. All rights reserved.
Conflict of interest statement
Competing Interest Statement A.P.S. holds stock in Krystal Biotech Inc. and consults for and has ownership interests in Zikani Therapeutics.
Figures


References
-
- Christiano AM, Anhalt G, Gibbons S, Bauer EA, Uitto J, Premature termination codons in the type VII collagen gene (COL7A1) underlie severe, mutilating recessive dystrophic epidermolysis bullosa, Genomics 21 (1) (1994) 160–168. - PubMed
-
- Fine JD, Bruckner-Tuderman L, Eady RA, Bauer EA, Bauer JW, Has C, Heagerty A, Hintner H, Hovnanian A, Jonkman MF, Leigh I, Marinkovich MP, Martinez AE, McGrath JA, Mellerio JE, Moss C, Murrell DF, Shimizu H, Uitto J, Woodley D, Zambruno G, Inherited epidermolysis bullosa: updated recommendations on diagnosis and classification, J Am Acad Dermatol 70 (6) (2014) 1103–1126. - PubMed
-
- Fine JD, Johnson LB, Weiner M, Li KP, Suchindran C, Epidermolysis bullosa and the risk of life-threatening cancers: the National EB Registry experience, 1986-2006, J Am Acad Dermatol 60 (2) (2009) 203–211. - PubMed
-
- Burgeson RE, Type VII collagen, anchoring fibrils, and epidermolysis bullosa, J Invest Dermatol 101 (3) (1993) 252–255. - PubMed
-
- Atanasova VS, Russell RJ, Webster TG, Cao Q, Agarwal P, Lim YZ, Krishnan S, Fuentes I, Guttmann-Gruber C, McGrath JA, Salas-Alanis JC, Fertala A, South AP, Thrombospondin-1 Is a Major Activator of TGF-beta Signaling in Recessive Dystrophic Epidermolysis Bullosa Fibroblasts, J Invest Dermatol 139 (7) (2019) 1497–1505 e5. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Miscellaneous

