Comparative genomic analyses of multiple backcross mouse populations suggest SGCG as a novel potential obesity-modifier gene
- PMID: 35796564
- PMCID: PMC9703946
- DOI: 10.1093/hmg/ddac150
Comparative genomic analyses of multiple backcross mouse populations suggest SGCG as a novel potential obesity-modifier gene
Abstract
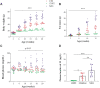
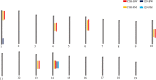
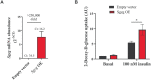
To nominate novel disease genes for obesity and type 2 diabetes (T2D), we recently generated two mouse backcross populations of the T2D-susceptible New Zealand Obese (NZO/HI) mouse strain and two genetically different, lean and T2D-resistant strains, 129P2/OlaHsd and C3HeB/FeJ. Comparative linkage analysis of our two female backcross populations identified seven novel body fat-associated quantitative trait loci (QTL). Only the locus Nbw14 (NZO body weight on chromosome 14) showed linkage to obesity-related traits in both backcross populations, indicating that the causal gene variant is likely specific for the NZO strain as NZO allele carriers in both crosses displayed elevated body weight and fat mass. To identify candidate genes for Nbw14, we used a combined approach of gene expression and haplotype analysis to filter for NZO-specific gene variants in gonadal white adipose tissue, defined as the main QTL-target tissue. Only two genes, Arl11 and Sgcg, fulfilled our candidate criteria. In addition, expression QTL analysis revealed cis-signals for both genes within the Nbw14 locus. Moreover, retroviral overexpression of Sgcg in 3T3-L1 adipocytes resulted in increased insulin-stimulated glucose uptake. In humans, mRNA levels of SGCG correlated with body mass index and body fat mass exclusively in diabetic subjects, suggesting that SGCG may present a novel marker for metabolically unhealthy obesity. In conclusion, our comparative-cross analysis could substantially improve the mapping resolution of the obesity locus Nbw14. Future studies will throw light on the mechanism by which Sgcg may protect from the development of obesity.
© The Author(s) 2022. Published by Oxford University Press.
Figures






References
-
- Centers for Disease Control and Prevention (2020) National Diabetes Statistics Report website. https://www.cdc.gov/diabetes/data/statistics-report/index.html.
-
- Mahajan, A., Taliun, D., Thurner, M., Robertson, N.R., Torres, J.M., Rayner, N.W., Payne, A.J., Steinthorsdottir, V., Scott, R.A., Grarup, N. et al. (2018) Fine-mapping type 2 diabetes loci to single-variant resolution using high-density imputation and islet-specific epigenome maps. Nat. Genet., 50, 1505–1513. - PMC - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Medical
Molecular Biology Databases
Research Materials

