The landscape of enhancer RNA identify prognosis-related molecular subtypes in gastric cancer
- PMID: 35801342
- PMCID: PMC9883414
- DOI: 10.1002/cam4.4959
The landscape of enhancer RNA identify prognosis-related molecular subtypes in gastric cancer
Abstract
Background: Enhancer RNAs (eRNAs), the transcriptional products of active enhancers, are of great significance in the initial progression of cancers. However, the biological function and bioinformatics profiles of eRNA in gastric cancer remains largely enigmatic.
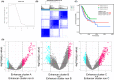
Methods: Firstly, STAD were clustered into three subtypes with the data of eRNA expression from TCeA. Then we explored the difference of the tumor immune microenvironment, transcription levels, and transcription regulation among the three clusters. Finally, samples collected from 12 patients diagnosed with STAD were used to conduct qRT-PCR, verifying the conclusion based on network database.
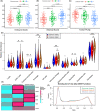
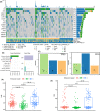
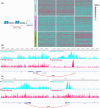
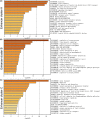
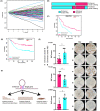
Results: The three clusters were detected to have different tumor microenvironments: Cluster A has an immune "cold" microenvironment. While cluster B features as more infiltration of immune cells, accompanied with higher expression of immune checkpoints such as PDCD1, LAG3, and TIGIT. Besides, Cluster C shows a higher stromal feature with B lineage, neutrophils, and fibroblasts. Further analyses indicated that CpG island methylation level of Cluster B is different from the other two clusters. Meanwhile, Cluster A and B showed significant enrichment of TP53 and KRAS mutation respectively while Cluster C has higher tumor mutation burden (TMB) and microsatellite instability (MSI). With the elaboration of transcriptional regulation of epigenetic clustering, we detected that Cluster A enriched in epithelial phenotype pathways. Cluster B enriched in cell-cell adhesion. Cluster C enriched in fibroblast proliferation. The clinical cohort show that Cluster B patients have lower interstitial cell characteristics and CAF infiltration.
Conclusion: We identified three unique epigenetic clusters of STAD through the differential activation of super-enhancers, and identified Cluster B with a higher immune infiltrating and a better prognosis, which provides a novel understanding of eRNAs and potential clinical applicability of eRNA-based molecular subtypes in gastric cancer.
Keywords: eRNAs; gastric cancer; immune microenvironment; methylation; mutation.
© 2022 Haian Hospital. Cancer Medicine published by John Wiley & Sons Ltd.
Conflict of interest statement
The authors declared no conflicts of interest in this work.
Figures
Similar articles
-
Comprehensive analysis of tumor immune microenvironment and prognosis of m6A-related lncRNAs in gastric cancer.BMC Cancer. 2022 Mar 24;22(1):316. doi: 10.1186/s12885-022-09377-8. BMC Cancer. 2022. PMID: 35331183 Free PMC article.
-
An immunogenic cell death-related signature predicts prognosis and immunotherapy response in stomach adenocarcinoma.Apoptosis. 2023 Dec;28(11-12):1564-1583. doi: 10.1007/s10495-023-01879-5. Epub 2023 Aug 14. Apoptosis. 2023. PMID: 37580435
-
Identification of cuproptosis-related subtypes, construction of a prognosis model, and tumor microenvironment landscape in gastric cancer.Front Immunol. 2022 Nov 21;13:1056932. doi: 10.3389/fimmu.2022.1056932. eCollection 2022. Front Immunol. 2022. PMID: 36479114 Free PMC article.
-
Identification and characterization of nucleotide metabolism and neuroendocrine regulation-associated modification patterns in stomach adenocarcinoma with auxiliary prognostic assessment and immunotherapy response prediction.Front Endocrinol (Lausanne). 2023 Jan 16;13:1076521. doi: 10.3389/fendo.2022.1076521. eCollection 2022. Front Endocrinol (Lausanne). 2023. PMID: 36726460 Free PMC article. Review.
-
A novel necroptosis-related gene index for predicting prognosis and a cold tumor immune microenvironment in stomach adenocarcinoma.Front Immunol. 2022 Oct 27;13:968165. doi: 10.3389/fimmu.2022.968165. eCollection 2022. Front Immunol. 2022. PMID: 36389725 Free PMC article. Review.
Cited by
-
Comprehensive Multiomics Analyses Establish the Optimal Prognostic Model for Resectable Gastric Cancer : Prognosis Prediction for Resectable GC.Ann Surg Oncol. 2024 Mar;31(3):2078-2089. doi: 10.1245/s10434-023-14249-x. Epub 2023 Nov 23. Ann Surg Oncol. 2024. PMID: 37996637
References
-
- Nelen SD, Bosscha K, Lemmens V, Hartgrink HH, Verhoeven RHA, de Wilt JHW. Morbidity and mortality according to age following gastrectomy for gastric cancer. Br J Surg. 2018;105(9):1163‐1170. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Medical
Research Materials
Miscellaneous

