Highly efficient CRISPR-mediated large DNA docking and multiplexed prime editing using a single baculovirus
- PMID: 35801912
- PMCID: PMC9303279
- DOI: 10.1093/nar/gkac587
Highly efficient CRISPR-mediated large DNA docking and multiplexed prime editing using a single baculovirus
Abstract
CRISPR-based precise gene-editing requires simultaneous delivery of multiple components into living cells, rapidly exceeding the cargo capacity of traditional viral vector systems. This challenge represents a major roadblock to genome engineering applications. Here we exploit the unmatched heterologous DNA cargo capacity of baculovirus to resolve this bottleneck in human cells. By encoding Cas9, sgRNA and Donor DNAs on a single, rapidly assembled baculoviral vector, we achieve with up to 30% efficacy whole-exon replacement in the intronic β-actin (ACTB) locus, including site-specific docking of very large DNA payloads. We use our approach to rescue wild-type podocin expression in steroid-resistant nephrotic syndrome (SRNS) patient derived podocytes. We demonstrate single baculovirus vectored delivery of single and multiplexed prime-editing toolkits, achieving up to 100% cleavage-free DNA search-and-replace interventions without detectable indels. Taken together, we provide a versatile delivery platform for single base to multi-gene level genome interventions, addressing the currently unmet need for a powerful delivery system accommodating current and future CRISPR technologies without the burden of limited cargo capacity.
© The Author(s) 2022. Published by Oxford University Press on behalf of Nucleic Acids Research.
Figures

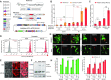
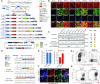
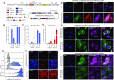


Similar articles
-
Assembly of Baculovirus Vectors for Multiplexed Prime Editing.Methods Mol Biol. 2024;2829:301-327. doi: 10.1007/978-1-0716-3961-0_24. Methods Mol Biol. 2024. PMID: 38951346
-
An engineered baculoviral protein and DNA co-delivery system for CRISPR-based mammalian genome editing.Nucleic Acids Res. 2024 Apr 12;52(6):3450-3468. doi: 10.1093/nar/gkae142. Nucleic Acids Res. 2024. PMID: 38412306 Free PMC article.
-
Baculoviral delivery of CRISPR/Cas9 facilitates efficient genome editing in human cells.PLoS One. 2017 Jun 22;12(6):e0179514. doi: 10.1371/journal.pone.0179514. eCollection 2017. PLoS One. 2017. PMID: 28640891 Free PMC article.
-
Lentiviral Vectors for Delivery of Gene-Editing Systems Based on CRISPR/Cas: Current State and Perspectives.Viruses. 2021 Jul 1;13(7):1288. doi: 10.3390/v13071288. Viruses. 2021. PMID: 34372494 Free PMC article. Review.
-
Pre-clinical non-viral vectors exploited for in vivo CRISPR/Cas9 gene editing: an overview.Biomater Sci. 2022 Jun 28;10(13):3410-3432. doi: 10.1039/d1bm01452h. Biomater Sci. 2022. PMID: 35604372 Review.
Cited by
-
Evaluation of Baculoviruses as Gene Therapy Vectors for Brain Cancer.Viruses. 2023 Feb 22;15(3):608. doi: 10.3390/v15030608. Viruses. 2023. PMID: 36992317 Free PMC article.
-
Integrating Prime Editing and Cellular Reprogramming as Novel Strategies for Genetic Cardiac Disease Modeling and Treatment.Curr Cardiol Rep. 2024 Nov;26(11):1197-1208. doi: 10.1007/s11886-024-02118-2. Epub 2024 Sep 11. Curr Cardiol Rep. 2024. PMID: 39259489 Free PMC article. Review.
-
Clinical and Translational Landscape of Viral Gene Therapies.Cells. 2024 Nov 19;13(22):1916. doi: 10.3390/cells13221916. Cells. 2024. PMID: 39594663 Free PMC article. Review.
-
A naked-eye biosensing system based on one-pot RPA-CRISPR/Cas12a driver G4-hemin self-assembly for Mycobacterium tuberculosis.Front Chem. 2025 Aug 7;13:1631086. doi: 10.3389/fchem.2025.1631086. eCollection 2025. Front Chem. 2025. PMID: 40851838 Free PMC article.
-
Tuning VSV-G Expression Improves Baculovirus Integrity, Stability and Mammalian Cell Transduction Efficiency.Viruses. 2024 Sep 17;16(9):1475. doi: 10.3390/v16091475. Viruses. 2024. PMID: 39339951 Free PMC article.
References
Publication types
MeSH terms
Substances
Associated data
Grants and funding
LinkOut - more resources
Full Text Sources
Research Materials
Miscellaneous

