Single-cell transcriptome atlas reveals developmental trajectories and a novel metabolic pathway of catechin esters in tea leaves
- PMID: 35810348
- PMCID: PMC9616531
- DOI: 10.1111/pbi.13891
Single-cell transcriptome atlas reveals developmental trajectories and a novel metabolic pathway of catechin esters in tea leaves
Abstract
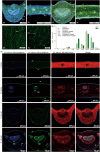
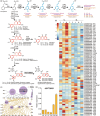
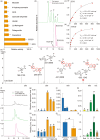
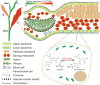
The tea plant is an economically important woody beverage crop. The unique taste of tea is evoked by certain metabolites, especially catechin esters, whereas their precise formation mechanism in different cell types remains unclear. Here, a fast protoplast isolation method was established and the transcriptional profiles of 16 977 single cells from 1st and 3rd leaves were investigated. We first identified 79 marker genes based on six isolated tissues and constructed a transcriptome atlas, mapped developmental trajectories and further delineated the distribution of different cell types during leaf differentiation and genes associated with cell fate transformation. Interestingly, eight differently expressed genes were found to co-exist at four branch points. Genes involved in the biosynthesis of certain metabolites showed cell- and development-specific characteristics. An unexpected catechin ester glycosyltransferase was characterized for the first time in plants by a gene co-expression network in mesophyll cells. Thus, the first single-cell transcriptional landscape in woody crop leave was reported and a novel metabolism pathway of catechin esters in plants was discovered.
Keywords: ScRNA-seq; catechin; development; glycosyltransferase; leaf; tea plant.
© 2022 The Authors. Plant Biotechnology Journal published by Society for Experimental Biology and The Association of Applied Biologists and John Wiley & Sons Ltd.
Conflict of interest statement
The authors declare no competing interests.
Figures




References
-
- Bonawitz, N.D. and Chapple, C. (2010) The genetics of lignin biosynthesis: connecting genotype to phenotype. Annu. Rev. Genet. 44, 337–363. - PubMed
-
- Boudolf, V. , Rombauts, S. , Naudts, M. , Inzé, D. and De Veylder, L. (2001) Identification of novel cyclin‐dependent kinases interacting with the CKS1 protein of Arabidopsis . J. Exp. Bot. 52, 1381–1382. - PubMed
-
- Butt, A.D. (1985) A general method for the high‐yield isolation of mesophyll protoplasts from deciduous tree species. Plant Sci. 42, 55–59.
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources

