Decoding RNA Editing Sites Through Transcriptome Analysis in Rice Under Alkaline Stress
- PMID: 35812946
- PMCID: PMC9260663
- DOI: 10.3389/fpls.2022.892729
Decoding RNA Editing Sites Through Transcriptome Analysis in Rice Under Alkaline Stress
Abstract
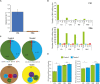
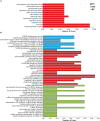
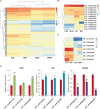
Ribonucleic acid editing (RE) is a post-transcriptional process that altered the genetics of RNA which provide the extra level of gene expression through insertion, deletions, and substitutions. In animals, it converts nucleotide residues C-U. Similarly in plants, the role of RNA editing sites (RES) in rice under alkaline stress is not fully studied. Rice is a staple food for most of the world population. Alkaline stress cause reduction in yield. Here, we explored the effect of alkaline stress on RES in the whole mRNA from rice chloroplast and mitochondria. Ribonucleic acid editing sites in both genomes (3336 RESs) including chloroplast (345 RESs) and mitochondria (2991 RESs) with average RES efficiency ∼55% were predicted. Our findings showed that majority of editing events found in non-synonymous codon changes and change trend in amino acids was hydrophobic. Four types of RNA editing A-G (A-I), C-T (C-U), G-A, and T-C were identified in treated and untreated samples. Overall, RNA editing efficiency was increased in the treated samples. Analysis of Gene Ontology revealed that mapped genes were engaged in many biological functions and molecular processes. We also checked the expression of pentatricopeptide repeat (PPR), organelle zinc-finger (OZI), and multiple organellar RNA editing factors/RNA editing factor interacting proteins genes in control and treatment, results revealed upregulation of PPR and OZ1 genes in treated samples. This induction showed the role of these genes in RNA editing. The current findings report that RNA editing increased under alkaline stress which may contribute in adaptation for rice by changing amino acids in edited genes (88 genes). These findings will provide basis for identification of RES in other crops and also will be useful in alkaline tolerance development in rice.
Keywords: MORF; OZ1; PPR; RNA editing; RNA-seq; alkaline stress; rice.
Copyright © 2022 Rehman, Uzair, Chao, Khan and Chen.
Conflict of interest statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Figures

References
-
- Andrews S. (2010). FastQC: A Quality Control Tool for High Throughput Sequence Data. Cambridge: Babraham Bioinformatics, Babraham Institute.
-
- Benjamini Y., Hochberg Y. (1995). Controlling the false discovery rate: a practical and powerful approach to multiple testing. J. R. Stat. Soc. 57 289–300.
LinkOut - more resources
Full Text Sources

