Malin restoration as proof of concept for gene therapy for Lafora disease
- PMID: 35813879
- PMCID: PMC9260307
- DOI: 10.1093/braincomms/fcac168
Malin restoration as proof of concept for gene therapy for Lafora disease
Abstract
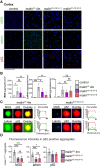
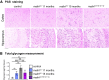
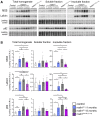
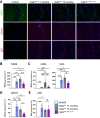
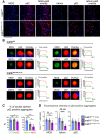
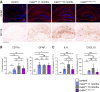
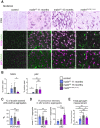
Lafora disease is a fatal neurodegenerative childhood dementia caused by loss-of-function mutations in either the laforin or malin gene. The hallmark of the disease is the accumulation of abnormal glycogen aggregates known as Lafora bodies (LBs) in the brain and other tissues. These aggregates are responsible for the pathological features of the disease. As a monogenic disorder, Lafora disease is a good candidate for gene therapy-based approaches. However, most patients are diagnosed after the appearance of the first symptoms and thus when LBs are already present in the brain. In this context, it was not clear whether the restoration of a normal copy of the defective gene (either laforin or malin) would prove effective. Here we evaluated the effect of restoring malin in a malin-deficient mouse model of Lafora disease as a proof of concept for gene replacement therapy. To this end, we generated a malin-deficient mouse in which malin expression can be induced at a certain time. Our results reveal that malin restoration at an advanced stage of the disease arrests the accumulation of LBs in brain and muscle, induces the degradation of laforin and glycogen synthase bound to the aggregates, and ameliorates neuroinflammation. These results identify malin restoration as the first therapeutic strategy to show effectiveness when applied at advanced stages of Lafora disease.
Keywords: Lafora disease; gene therapy; glycogen; neurodegeneration; neuroinflammation.
© The Author(s) 2022. Published by Oxford University Press on behalf of the Guarantors of Brain.
Figures


Similar articles
-
Therapeutic Window for the Treatment of Lafora Disease.In: Noebels JL, Avoli M, Rogawski MA, Vezzani A, Delgado-Escueta AV, editors. Jasper's Basic Mechanisms of the Epilepsies. 5th edition. New York: Oxford University Press; 2024. Chapter 53. In: Noebels JL, Avoli M, Rogawski MA, Vezzani A, Delgado-Escueta AV, editors. Jasper's Basic Mechanisms of the Epilepsies. 5th edition. New York: Oxford University Press; 2024. Chapter 53. PMID: 39637182 Free Books & Documents. Review.
-
Lafora disease: Current biology and therapeutic approaches.Rev Neurol (Paris). 2022 Apr;178(4):315-325. doi: 10.1016/j.neurol.2021.06.006. Epub 2021 Jul 21. Rev Neurol (Paris). 2022. PMID: 34301405 Free PMC article. Review.
-
Increased laforin and laforin binding to glycogen underlie Lafora body formation in malin-deficient Lafora disease.J Biol Chem. 2012 Jul 20;287(30):25650-9. doi: 10.1074/jbc.M111.331611. Epub 2012 Jun 5. J Biol Chem. 2012. PMID: 22669944 Free PMC article.
-
Lafora Disease: A Ubiquitination-Related Pathology.Cells. 2018 Jul 26;7(8):87. doi: 10.3390/cells7080087. Cells. 2018. PMID: 30050012 Free PMC article. Review.
-
Abnormal glycogen chain length pattern, not hyperphosphorylation, is critical in Lafora disease.EMBO Mol Med. 2017 Jul;9(7):906-917. doi: 10.15252/emmm.201707608. EMBO Mol Med. 2017. PMID: 28536304 Free PMC article.
Cited by
-
Role of Astrocytes in the Pathophysiology of Lafora Disease and Other Glycogen Storage Disorders.Cells. 2023 Feb 24;12(5):722. doi: 10.3390/cells12050722. Cells. 2023. PMID: 36899857 Free PMC article. Review.
-
Neurological glycogen storage diseases and emerging therapeutics.Neurotherapeutics. 2024 Sep;21(5):e00446. doi: 10.1016/j.neurot.2024.e00446. Epub 2024 Sep 14. Neurotherapeutics. 2024. PMID: 39277505 Free PMC article. Review.
-
Glycogen accumulation modulates life span in a mouse model of amyotrophic lateral sclerosis.J Neurochem. 2024 May;168(5):744-759. doi: 10.1111/jnc.15906. Epub 2023 Jul 4. J Neurochem. 2024. PMID: 37401737 Free PMC article.
-
Lafora Disease: A Case Report and Evolving Treatment Advancements.Brain Sci. 2023 Dec 6;13(12):1679. doi: 10.3390/brainsci13121679. Brain Sci. 2023. PMID: 38137127 Free PMC article.
-
Empagliflozin Repurposing for Lafora Disease: A Pilot Clinical Trial and Preclinical Investigation of Novel Therapeutic Targets.Methods Protoc. 2025 May 6;8(3):48. doi: 10.3390/mps8030048. Methods Protoc. 2025. PMID: 40407475 Free PMC article.
References
Grants and funding
LinkOut - more resources
Full Text Sources
Research Materials
