Class I Polyhydroxyalkanoate (PHA) Synthase Increased Polylactic Acid Production in Engineered Escherichia Coli
- PMID: 35814019
- PMCID: PMC9261260
- DOI: 10.3389/fbioe.2022.919969
Class I Polyhydroxyalkanoate (PHA) Synthase Increased Polylactic Acid Production in Engineered Escherichia Coli
Abstract
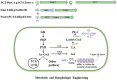
Polylactic acid (PLA), a homopolymer of lactic acid (LA), is a bio-derived, biocompatible, and biodegradable polyester. The evolved class II PHA synthase (PhaC1 Ps6-19) was commonly utilized in the de novo biosynthesis of PLA from biomass. This study tested alternative class I PHA synthase (PhaC Cs ) from Chromobacterium sp. USM2 in engineered Escherichia coli for the de novo biosynthesis of PLA from glucose. The results indicated that PhaC Cs had better performance in PLA production than that of class II synthase PhaC1 Ps6-19. In addition, the sulA gene was engineered in PLA-producing strains for morphological engineering. The morphologically engineered strains present increased PLA production. This study also tested fused propionyl-CoA transferase and lactate dehydrogenase A (fused Pct Cp /LdhA) in engineered E. coli and found that fused Pct Cp /LdhA did not apparently improve the PLA production. After systematic engineering, the highest PLA production was achieved by E. coli MS6 (with PhaC Cs and sulA), which could produce up to 955.0 mg/L of PLA in fed-batch fermentation with the cell dry weights of 2.23%, and the average molecular weight of produced PLA could reach 21,000 Da.
Keywords: biopolyester; class I polyhydroxyalkanoate synthase; degradable polymer; engineered E. coli; polylactic acid.
Copyright © 2022 Shi, Li, Yang, Miao, Yang, Pandhal and Zou.
Conflict of interest statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Figures


References
LinkOut - more resources
Full Text Sources
Miscellaneous

