Spatial and Transcriptomic Analysis of Perineural Invasion in Oral Cancer
- PMID: 35819260
- PMCID: PMC9560986
- DOI: 10.1158/1078-0432.CCR-21-4543
Spatial and Transcriptomic Analysis of Perineural Invasion in Oral Cancer
Abstract
Purpose: Perineural invasion (PNI), a common occurrence in oral squamous cell carcinomas, is associated with poor survival. Consequently, these tumors are treated aggressively. However, diagnostic criteria of PNI vary and its role as an independent predictor of prognosis has not been established. To address these knowledge gaps, we investigated spatial and transcriptomic profiles of PNI-positive and PNI-negative nerves.
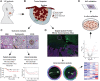
Experimental design: Tissue sections from 142 patients were stained with S100 and cytokeratin antibodies. Nerves were identified in two distinct areas: tumor bulk and margin. Nerve diameter and nerve-to-tumor distance were assessed; survival analyses were performed. Spatial transcriptomic analysis of nerves at varying distances from tumor was performed with NanoString GeoMx Digital Spatial Profiler Transcriptomic Atlas.
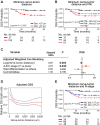
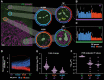
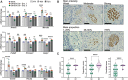
Results: PNI is an independent predictor of poor prognosis among patients with metastasis-free lymph nodes. Patients with close nerve-tumor distance have poor outcomes even if diagnosed as PNI negative using current criteria. Patients with large nerve(s) in the tumor bulk survive poorly, suggesting that even PNI-negative nerves facilitate tumor progression. Diagnostic criteria were supported by spatial transcriptomic analyses of >18,000 genes; nerves in proximity to cancer exhibit stress and growth response changes that diminish with increasing nerve-tumor distance. These findings were validated in vitro and in human tissue.
Conclusions: This is the first study in human cancer with high-throughput gene expression analysis in nerves with striking correlations between transcriptomic profile and clinical outcomes. Our work illuminates nerve-cancer interactions suggesting that cancer-induced injury modulates neuritogenesis, and supports reclassification of PNI based on nerve-tumor distance rather than current subjective criteria.
©2022 The Authors; Published by the American Association for Cancer Research.
Figures



Comment in
- 1078-0432. doi: 10.1158/1078-0432.CCR-28-16-HI
Similar articles
-
Subcategorization of Perineural Invasion and Its Impact on Survival in Patients with Oral Squamous Cell Carcinoma.Head Neck Pathol. 2023 Jun;17(2):383-392. doi: 10.1007/s12105-022-01512-y. Epub 2022 Dec 8. Head Neck Pathol. 2023. PMID: 36480091 Free PMC article.
-
Quantitative Measurement of Perineural Invasion for Prognosis Analysis of Oral Cavity Cancer Treated by Radical Surgery With or Without Adjuvant Therapy.Technol Cancer Res Treat. 2023 Jan-Dec;22:15330338231176366. doi: 10.1177/15330338231176366. Technol Cancer Res Treat. 2023. PMID: 37264638 Free PMC article.
-
Perineural invasion in squamous cell carcinoma of the head and neck.Arch Otolaryngol Head Neck Surg. 1998 Jun;124(6):637-40. doi: 10.1001/archotol.124.6.637. Arch Otolaryngol Head Neck Surg. 1998. PMID: 9639472
-
Perineural invasion-associated biomarkers for tumor development.Biomed Pharmacother. 2022 Nov;155:113691. doi: 10.1016/j.biopha.2022.113691. Epub 2022 Sep 12. Biomed Pharmacother. 2022. PMID: 36095958 Review.
-
The 3D's of Neural Phenotypes in Oral Cancer: Distance, Diameter, and Density.Adv Biol (Weinh). 2023 Feb;7(2):e2200188. doi: 10.1002/adbi.202200188. Epub 2022 Nov 14. Adv Biol (Weinh). 2023. PMID: 36373694 Free PMC article. Review.
Cited by
-
[Advances of spatial omics in the individualized diagnosis and treatment of head and neck cancer].Lin Chuang Er Bi Yan Hou Tou Jing Wai Ke Za Zhi. 2023 Sep;37(9):729-733;739. doi: 10.13201/j.issn.2096-7993.2023.09.008. Lin Chuang Er Bi Yan Hou Tou Jing Wai Ke Za Zhi. 2023. PMID: 37830120 Free PMC article. Chinese.
-
Investigation of the correlation between AGRN expression and perineural invasion in colon cancer.Front Mol Biosci. 2024 Dec 3;11:1510478. doi: 10.3389/fmolb.2024.1510478. eCollection 2024. Front Mol Biosci. 2024. PMID: 39691475 Free PMC article.
-
Tumor Neurobiology in the Pathogenesis and Therapy of Head and Neck Cancer.Cells. 2024 Jan 30;13(3):256. doi: 10.3390/cells13030256. Cells. 2024. PMID: 38334648 Free PMC article. Review.
-
Nerves at Play: The Peripheral Nervous System in Extracranial Malignancies.Cancer Discov. 2025 Jan 13;15(1):52-68. doi: 10.1158/2159-8290.CD-23-0397. Cancer Discov. 2025. PMID: 39801235 Free PMC article. Review.
-
Mapping cancer biology in space: applications and perspectives on spatial omics for oncology.Mol Cancer. 2024 Jan 30;23(1):26. doi: 10.1186/s12943-024-01941-z. Mol Cancer. 2024. PMID: 38291400 Free PMC article. Review.
References
-
- Zhao B, Lv W, Mei D, Luo R, Bao S, Huang B, et al. . Perineural invasion as a predictive factor for survival outcome in gastric cancer patients: a systematic review and meta-analysis. J Clin Pathol 2020;73:544–51. - PubMed
-
- Liebig C, Ayala G, Wilks JA, Berger DH, Albo D. Perineural invasion in cancer: a review of the literature. Cancer 2009;115:3379–91. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Medical

