Histones of Neutrophil Extracellular Traps Induce CD11b Expression in Brain Pericytes Via Dectin-1 after Traumatic Brain Injury
- PMID: 35819574
- PMCID: PMC9554061
- DOI: 10.1007/s12264-022-00902-0
Histones of Neutrophil Extracellular Traps Induce CD11b Expression in Brain Pericytes Via Dectin-1 after Traumatic Brain Injury
Abstract
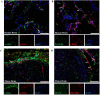
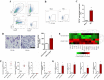
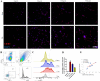
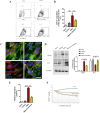
The brain pericyte is a unique and indispensable part of the blood-brain barrier (BBB), and contributes to several pathological processes in traumatic brain injury (TBI). However, the cellular and molecular mechanisms by which pericytes are regulated in the damaged brain are largely unknown. Here, we show that the formation of neutrophil extracellular traps (NETs) induces the appearance of CD11b+ pericytes after TBI. These CD11b+ pericyte subsets are characterized by increased permeability and pro-inflammatory profiles compared to CD11b- pericytes. Moreover, histones from NETs by Dectin-1 facilitate CD11b induction in brain pericytes in PKC-c-Jun dependent manner, resulting in neuroinflammation and BBB dysfunction after TBI. These data indicate that neutrophil-NET-pericyte and histone-Dectin-1-CD11b are possible mechanisms for the activation and dysfunction of pericytes. Targeting NETs formation and Dectin-1 are promising means of treating TBI.
Keywords: Dectin-1; NET; Neutrophil; Pericyte; TBI.
© 2022. Center for Excellence in Brain Science and Intelligence Technology, Chinese Academy of Sciences.
Conflict of interest statement
The authors all declare that they have no competing interests.
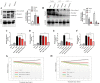
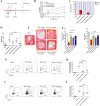
Figures


Similar articles
-
Blood-Brain Barrier (BBB) Dysfunction in CNS Diseases: Paying Attention to Pericytes.CNS Neurosci Ther. 2025 May;31(5):e70422. doi: 10.1111/cns.70422. CNS Neurosci Ther. 2025. PMID: 40371544 Free PMC article. Review.
-
Impairment of pericyte-endothelium crosstalk leads to blood-brain barrier dysfunction following traumatic brain injury.Exp Neurol. 2019 Jul;317:260-270. doi: 10.1016/j.expneurol.2019.03.014. Epub 2019 Mar 26. Exp Neurol. 2019. PMID: 30926390
-
RAGE mediates hippocampal pericyte responses and neurovascular unit lesions after TBI.Exp Neurol. 2024 Oct;380:114912. doi: 10.1016/j.expneurol.2024.114912. Epub 2024 Aug 2. Exp Neurol. 2024. PMID: 39097075
-
Inhibition of neutrophil extracellular trap formation attenuates NLRP1-dependent neuronal pyroptosis via STING/IRE1α pathway after traumatic brain injury in mice.Front Immunol. 2023 Apr 14;14:1125759. doi: 10.3389/fimmu.2023.1125759. eCollection 2023. Front Immunol. 2023. PMID: 37143681 Free PMC article.
-
Pericyte Loss in Diseases.Cells. 2023 Jul 26;12(15):1931. doi: 10.3390/cells12151931. Cells. 2023. PMID: 37566011 Free PMC article. Review.
Cited by
-
S100A9 as a potential novel target for experimental autoimmune cystitis and interstitial cystitis/bladder pain syndrome.Biomark Res. 2025 May 9;13(1):72. doi: 10.1186/s40364-025-00763-5. Biomark Res. 2025. PMID: 40346703 Free PMC article.
-
FOXO1 reshapes neutrophils to aggravate acute brain damage and promote late depression after traumatic brain injury.Mil Med Res. 2024 Mar 31;11(1):20. doi: 10.1186/s40779-024-00523-w. Mil Med Res. 2024. PMID: 38556884 Free PMC article.
-
Neutrophil Extracellular Traps Regulate Surgical Brain Injury by Activating the cGAS-STING Pathway.Cell Mol Neurobiol. 2024 Apr 18;44(1):36. doi: 10.1007/s10571-024-01470-9. Cell Mol Neurobiol. 2024. PMID: 38637346 Free PMC article.
-
Neutrophil extracellular traps induced by neoadjuvant chemotherapy of breast cancer promotes vascular endothelial damage.Breast Cancer Res. 2025 Apr 23;27(1):61. doi: 10.1186/s13058-025-02011-y. Breast Cancer Res. 2025. PMID: 40270028 Free PMC article.
-
Blood-Brain Barrier (BBB) Dysfunction in CNS Diseases: Paying Attention to Pericytes.CNS Neurosci Ther. 2025 May;31(5):e70422. doi: 10.1111/cns.70422. CNS Neurosci Ther. 2025. PMID: 40371544 Free PMC article. Review.
References
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Medical
Research Materials
Miscellaneous

