RNANetMotif: Identifying sequence-structure RNA network motifs in RNA-protein binding sites
- PMID: 35819951
- PMCID: PMC9275694
- DOI: 10.1371/journal.pcbi.1010293
RNANetMotif: Identifying sequence-structure RNA network motifs in RNA-protein binding sites
Abstract
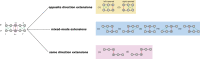
RNA molecules can adopt stable secondary and tertiary structures, which are essential in mediating physical interactions with other partners such as RNA binding proteins (RBPs) and in carrying out their cellular functions. In vivo and in vitro experiments such as RNAcompete and eCLIP have revealed in vitro binding preferences of RBPs to RNA oligomers and in vivo binding sites in cells. Analysis of these binding data showed that the structure properties of the RNAs in these binding sites are important determinants of the binding events; however, it has been a challenge to incorporate the structure information into an interpretable model. Here we describe a new approach, RNANetMotif, which takes predicted secondary structure of thousands of RNA sequences bound by an RBP as input and uses a graph theory approach to recognize enriched subgraphs. These enriched subgraphs are in essence shared sequence-structure elements that are important in RBP-RNA binding. To validate our approach, we performed RNA structure modeling via coarse-grained molecular dynamics folding simulations for selected 4 RBPs, and RNA-protein docking for LIN28B. The simulation results, e.g., solvent accessibility and energetics, further support the biological relevance of the discovered network subgraphs.
Conflict of interest statement
none
Figures




Similar articles
-
Finding RNA structure in the unstructured RBPome.BMC Genomics. 2018 Feb 20;19(1):154. doi: 10.1186/s12864-018-4540-1. BMC Genomics. 2018. PMID: 29463232 Free PMC article.
-
PRIESSTESS: interpretable, high-performing models of the sequence and structure preferences of RNA-binding proteins.Nucleic Acids Res. 2022 Oct 28;50(19):e111. doi: 10.1093/nar/gkac694. Nucleic Acids Res. 2022. PMID: 36018788 Free PMC article.
-
RNA-protein binding motifs mining with a new hybrid deep learning based cross-domain knowledge integration approach.BMC Bioinformatics. 2017 Feb 28;18(1):136. doi: 10.1186/s12859-017-1561-8. BMC Bioinformatics. 2017. PMID: 28245811 Free PMC article.
-
Finding the target sites of RNA-binding proteins.Wiley Interdiscip Rev RNA. 2014 Jan-Feb;5(1):111-30. doi: 10.1002/wrna.1201. Epub 2013 Nov 11. Wiley Interdiscip Rev RNA. 2014. PMID: 24217996 Free PMC article. Review.
-
Zooming in on protein-RNA interactions: a multi-level workflow to identify interaction partners.Biochem Soc Trans. 2020 Aug 28;48(4):1529-1543. doi: 10.1042/BST20191059. Biochem Soc Trans. 2020. PMID: 32820806 Free PMC article. Review.
Cited by
-
A weighted two-stage sequence alignment framework to identify motifs from ChIP-exo data.Patterns (N Y). 2024 Feb 2;5(3):100927. doi: 10.1016/j.patter.2024.100927. eCollection 2024 Mar 8. Patterns (N Y). 2024. PMID: 38487805 Free PMC article.
-
Discovery of RNA-Targeting Small Molecules: Challenges and Future Directions.MedComm (2020). 2025 Aug 24;6(9):e70342. doi: 10.1002/mco2.70342. eCollection 2025 Sep. MedComm (2020). 2025. PMID: 40859960 Free PMC article. Review.
-
Role of Optimization in RNA-Protein-Binding Prediction.Curr Issues Mol Biol. 2024 Feb 4;46(2):1360-1373. doi: 10.3390/cimb46020087. Curr Issues Mol Biol. 2024. PMID: 38392205 Free PMC article.
-
qNABpredict: Quick, accurate, and taxonomy-aware sequence-based prediction of content of nucleic acid binding amino acids.Protein Sci. 2023 Jan;32(1):e4544. doi: 10.1002/pro.4544. Protein Sci. 2023. PMID: 36519304 Free PMC article.
-
Inferring RNA-binding protein target preferences using adversarial domain adaptation.PLoS Comput Biol. 2022 Feb 24;18(2):e1009863. doi: 10.1371/journal.pcbi.1009863. eCollection 2022 Feb. PLoS Comput Biol. 2022. PMID: 35202389 Free PMC article.
References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources

