A super pan-genomic landscape of rice
- PMID: 35821092
- PMCID: PMC9525306
- DOI: 10.1038/s41422-022-00685-z
A super pan-genomic landscape of rice
Abstract
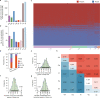
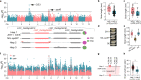
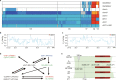
Pan-genomes from large natural populations can capture genetic diversity and reveal genomic complexity. Using de novo long-read assembly, we generated a graph-based super pan-genome of rice consisting of a 251-accession panel comprising both cultivated and wild species of Asian and African rice. Our pan-genome reveals extensive structural variations (SVs) and gene presence/absence variations. Additionally, our pan-genome enables the accurate identification of nucleotide-binding leucine-rich repeat genes and characterization of their inter- and intraspecific diversity. Moreover, we uncovered grain weight-associated SVs which specify traits by affecting the expression of their nearby genes. We characterized genetic variants associated with submergence tolerance, seed shattering and plant architecture and found independent selection for a common set of genes that drove adaptation and domestication in Asian and African rice. This super pan-genome facilitates pinpointing of lineage-specific haplotypes for trait-associated genes and provides insights into the evolutionary events that have shaped the genomic architecture of various rice species.
© 2022. The Author(s).
Conflict of interest statement
The authors declare no competing interests.
Figures




Comment in
-
The rice pangenome branches out.Cell Res. 2022 Oct;32(10):867-868. doi: 10.1038/s41422-022-00699-7. Cell Res. 2022. PMID: 35821091 Free PMC article. No abstract available.
References
-
- Gnanamanickam, S. S. Rice and its importance to human life. In: biological control of rice diseases. Progress in biological control. Springer, (2009).
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources

