Increased sensitivity of Aggregatibacter actinomycetemcomitans to human serum is mediated by induction of a bacteriophage
- PMID: 35833243
- PMCID: PMC10087258
- DOI: 10.1111/omi.12378
Increased sensitivity of Aggregatibacter actinomycetemcomitans to human serum is mediated by induction of a bacteriophage
Abstract
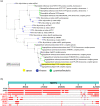
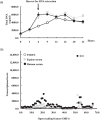
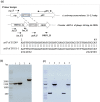
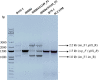
Aggregatibacter actinomycetemcomitans, a Gram-negative oral pathobiont causing aggressive periodontitis and systemic infections, demonstrates serum resistance. We have identified a dsDNA-tailed bacteriophage, S1249, which was found to convert from this microorganism inducible by human serum into a lytic state to kill the bacterium. This phage demonstrated active transcripts when exposed to human serum: 20% of genes were upregulated more than 10-fold, and 45% of them were upregulated 5-10-fold when the bacterium was grown in the presence of human serum compared to without the presence of human serum. Transcriptional activation when grown in equine serum was less pronounced. This phage demonstrated a tail with inner rigid tubes and an outer contractile sheath, features of Myoviridae spp. Further characterization revealed that the lysogenized integration of the phage in the chromosome of A. actinomycetemcomitans occurred between the genes encoding cold-shock DNA-binding domain-containing protein (csp) and glutamyl-tRNA synthetase (gltX). Both phage DNA integrated lysogeny and nonintegrated pseudolysogeny were identified in the infected bacterium. A newly generated, lysogenized strain using this phage displayed similar attributes, including 63% growth inhibition compared to its isogenic phage-free strain when in the presence of human serum. Our data suggest that bacteriophage S1249 can be induced in the presence of human serum and enters the lytic cycle, which reduces the viability of infected bacteria in vivo.
Keywords: Myoviridae; bacteriophages; gram-negative periodontal pathobionts; human serum; lytic cycle; pseudolysogeny.
© 2022 The Authors. Molecular Oral Microbiology published by John Wiley & Sons Ltd.
Conflict of interest statement
The authors declare no conflict of interest.
Figures



References
-
- Akrivopoulou, C. , Green, I. M. , Donos, N. , Nair, S. P. , & Ready, D. (2017). Aggregatibacter actinomycetemcomitans serotype prevalence and antibiotic resistance in a UK population with periodontitis. Journal of Global Antimicrobial Resistance, 10, 54–58. - PubMed
-
- Anton‐Vazquez, V. , Dworakowski, R. , Cannata, A. , Amin‐Youssef, G. , Gunning, M. , Papachristidis, A. , MacCarthy, P. , Baghai, M. , Deshpande, R. , Khan, H. , Byrne, J. , & Fife, A. (2022). 16S rDNA PCR for the aetiological diagnosis of culture‐negative infective endocarditis. Infection, 50(1), 243–249. 10.1007/s15010-021-01690-x - DOI - PubMed
-
- Ardila, C. M. , & Bedoya‐García, J. A. (2020). Antimicrobial resistance of Aggregatibacter actinomycetemcomitans, Porphyromonas gingivalis and Tannerella forsythia in periodontitis patients. Journal of Global Antimicrobial Resistance, 22, 215–218. - PubMed
-
- Asakawa, R. , Komatsuzawa, H. , Kawai, T. , Yamada, S. , Goncalves, R. B. , Izumi, S. , Fujiwara, T. , Nakano, Y. , Suzuki, N. , Uchida, Y. , Ouhara, K. , Shiba, H. , Taubman, M. A. , Kurihara, H. , & Sugai, M. (2003). Outer membrane protein 100, a versatile virulence factor of Actinobacillus actinomycetemcomitans . Molecular Microbiology, 50(4), 1125–1139. 10.1046/j.1365-2958.2003.03748.x - DOI - PubMed
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Miscellaneous

