Improved detection limits of J-coupled neurometabolites in the human brain at 7 T with a J-refocused sLASER sequence
- PMID: 35833462
- PMCID: PMC9788253
- DOI: 10.1002/nbm.4801
Improved detection limits of J-coupled neurometabolites in the human brain at 7 T with a J-refocused sLASER sequence
Abstract
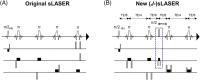
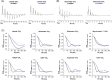
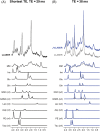
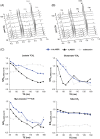
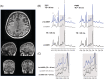
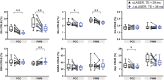
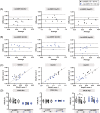
In a standard spin echo, the time evolution due to homonuclear couplings is not reversed, leading to echo time (TE)-dependent modulation of the signal amplitude and signal loss in the case of overlapping multiplet resonances. This has an adverse effect on quantification of several important metabolites such as glutamate and glutamine. Here, we propose a J-refocused variant of the sLASER sequence (J-sLASER) to improve quantification of J-coupled metabolites at ultrahigh field (UHF). The use of the sLASER sequence is particularly advantageous at UHF as it minimizes chemical shift displacement error and results in relatively homogenous refocusing. We simulated the MRS signal from brain metabolites over a broad range of TE values with sLASER and J-sLASER, and showed that the signal of J-coupled metabolites was increased with J-sLASER with TE values up to ~80 ms. We further simulated "brain-like" spectra with both sequences at the shortest TE available on our scanner. We showed that, despite the slightly longer TE, the J-sLASER sequence results in significantly lower Cramer-Rao lower bounds (CRLBs) for J-coupled metabolites compared with those obtained with sLASER. Following phantom validation, we acquired spectra from two brain regions in 10 healthy volunteers (age 38 ± 15 years) using both sequences. We showed that using J-sLASER results in a decrease of CRLBs for J-coupled metabolites. In particular, we measured a robust ~38% decrease in the mean CRLB (glutamine) in parietal white matter and posterior cingulate cortex (PCC). We further showed, in 10 additional healthy volunteers (age 34 ± 15 years), that metabolite quantification following two separate acquisitions with J-sLASER in the PCC was repeatable. The improvement in quantification of glutamine may in turn improve the independent quantification of glutamate, the main excitatory neurotransmitter in the brain, and will simultaneously help to track possible modulations of glutamine, which is a key player in the glutamatergic cycle in astrocytes.
Keywords: 7 T; J-coupled metabolites; J-refocused sLASER; aspartate; detection limits; glutamate; glutamine; human brain.
© 2022 The Authors. NMR in Biomedicine published by John Wiley & Sons Ltd.
Conflict of interest statement
The authors declare no conflicts of interest.
Figures

References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources

