Live-Cell Imaging of Sterculic Acid-a Naturally Occurring 1,2-Cyclopropene Fatty Acid-by Bioorthogonal Reaction with Turn-On Tetrazine-Fluorophore Conjugates
- PMID: 35838324
- PMCID: PMC9546306
- DOI: 10.1002/anie.202207640
Live-Cell Imaging of Sterculic Acid-a Naturally Occurring 1,2-Cyclopropene Fatty Acid-by Bioorthogonal Reaction with Turn-On Tetrazine-Fluorophore Conjugates
Abstract
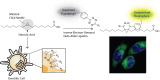
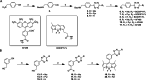
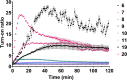
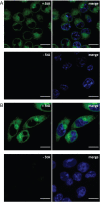
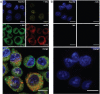
In the field of lipid research, bioorthogonal chemistry has made the study of lipid uptake and processing in living systems possible, whilst minimising biological properties arising from detectable pendant groups. To allow the study of unsaturated free fatty acids in live cells, we here report the use of sterculic acid, a 1,2-cyclopropene-containing oleic acid analogue, as a bioorthogonal probe. We show that this lipid can be readily taken up by dendritic cells without toxic side effects, and that it can subsequently be visualised using an inverse electron-demand Diels-Alder reaction with quenched tetrazine-fluorophore conjugates. In addition, the lipid can be used to identify changes in protein oleoylation after immune cell activation. Finally, this reaction can be integrated into a multiplexed bioorthogonal reaction workflow by combining it with two sequential copper-catalysed Huisgen ligation reactions. This allows for the study of multiple biomolecules in the cell simultaneously by multimodal confocal imaging.
Keywords: Click Chemistry; Cyclopropene; Inverse Electron-Demand Diels-Alder Reaction; Lipids; Sterculic Acid.
© 2022 The Authors. Angewandte Chemie International Edition published by Wiley-VCH GmbH.
Conflict of interest statement
The authors declare no conflict of interest.
Figures

Similar articles
-
Advances in Tetrazine Bioorthogonal Chemistry Driven by the Synthesis of Novel Tetrazines and Dienophiles.Acc Chem Res. 2018 May 15;51(5):1249-1259. doi: 10.1021/acs.accounts.8b00062. Epub 2018 Apr 11. Acc Chem Res. 2018. PMID: 29638113 Free PMC article.
-
Monochromophoric Design Strategy for Tetrazine-Based Colorful Bioorthogonal Probes with a Single Fluorescent Core Skeleton.J Am Chem Soc. 2018 Jan 24;140(3):974-983. doi: 10.1021/jacs.7b10433. Epub 2017 Dec 29. J Am Chem Soc. 2018. PMID: 29240995
-
Two-Photon and Multicolor Fluorogenic Bioorthogonal Probes Based on Tetrazine-Conjugated Naphthalene Fluorophores.Bioconjug Chem. 2020 May 20;31(5):1545-1550. doi: 10.1021/acs.bioconjchem.0c00197. Epub 2020 Apr 23. Bioconjug Chem. 2020. PMID: 32297734
-
Inverse Electron-Demand Diels-Alder Bioorthogonal Reactions.Top Curr Chem (Cham). 2016 Feb;374(1):3. doi: 10.1007/s41061-015-0005-z. Epub 2015 Dec 22. Top Curr Chem (Cham). 2016. PMID: 27572986 Review.
-
Photo-induced and Rapid Labeling of Tetrazine-Bearing Proteins via Cyclopropenone-Caged Bicyclononynes.Angew Chem Int Ed Engl. 2019 Oct 28;58(44):15876-15882. doi: 10.1002/anie.201908209. Epub 2019 Sep 24. Angew Chem Int Ed Engl. 2019. PMID: 31476269 Free PMC article. Review.
Cited by
-
Chemical Probes to Control and Visualize Lipid Metabolism in the Brain.Acc Chem Res. 2022 Nov 15;55(22):3205-3217. doi: 10.1021/acs.accounts.2c00521. Epub 2022 Oct 25. Acc Chem Res. 2022. PMID: 36283077 Free PMC article. Review.
-
Photoreactive bioorthogonal lipid probes and their applications in mammalian biology.RSC Chem Biol. 2022 Dec 12;4(1):37-46. doi: 10.1039/d2cb00174h. eCollection 2023 Jan 4. RSC Chem Biol. 2022. PMID: 36685253 Free PMC article.
-
Lipid Metabolism and Immune Function: Chemical Tools for Insights into T-Cell Biology.Chembiochem. 2025 Jun 3;26(11):e202400980. doi: 10.1002/cbic.202400980. Epub 2025 Apr 16. Chembiochem. 2025. PMID: 40162512 Free PMC article. Review.
-
A Lysosome-Targeted Tetrazine for Organelle-Specific Click-to-Release Chemistry in Antigen Presenting Cells.J Am Chem Soc. 2023 Jun 14;145(23):12630-12640. doi: 10.1021/jacs.3c02139. Epub 2023 Jun 3. J Am Chem Soc. 2023. PMID: 37269296 Free PMC article.
-
Click Chemistry: Reaction Rates and Their Suitability for Biomedical Applications.Bioconjug Chem. 2024 Jun 19;35(6):715-731. doi: 10.1021/acs.bioconjchem.4c00084. Epub 2024 May 22. Bioconjug Chem. 2024. PMID: 38775705 Free PMC article. Review.
References
-
- Wymann M. P., Schneiter R., Nat. Rev. Mol. Cell Biol. 2008, 9, 162–176. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources

