Optimizing Epitope Conformational Ensembles Using α-Synuclein Cyclic Peptide "Glycindel" Scaffolds: A Customized Immunogen Method for Generating Oligomer-Selective Antibodies for Parkinson's Disease
- PMID: 35840132
- PMCID: PMC9354795
- DOI: 10.1021/acschemneuro.1c00567
Optimizing Epitope Conformational Ensembles Using α-Synuclein Cyclic Peptide "Glycindel" Scaffolds: A Customized Immunogen Method for Generating Oligomer-Selective Antibodies for Parkinson's Disease
Abstract
Effectively presenting epitopes on immunogens, in order to raise conformationally selective antibodies through active immunization, is a central problem in treating protein misfolding diseases, particularly neurodegenerative diseases such as Alzheimer's disease or Parkinson's disease. We seek to selectively target conformations enriched in toxic, oligomeric propagating species while sparing the healthy forms of the protein that are often more abundant. To this end, we computationally modeled scaffolded epitopes in cyclic peptides by inserting/deleting a variable number of flanking glycines ("glycindels") to best mimic a misfolding-specific conformation of an epitope of α-synuclein enriched in the oligomer ensemble, as characterized by a region most readily disordered and solvent-exposed in a stressed, partially denatured protofibril. We screen and rank the cyclic peptide scaffolds of α-synuclein in silico based on their ensemble overlap properties with the fibril, oligomer-model and isolated monomer ensembles. We present experimental data of seeded aggregation that support nucleation rates consistent with computationally predicted cyclic peptide conformational similarity. We also introduce a method for screening against structured off-pathway targets in the human proteome by selecting scaffolds with minimal conformational similarity between their epitope and the same solvent-exposed primary sequence in structured human proteins. Different cyclic peptide scaffolds with variable numbers of glycines are predicted computationally to have markedly different conformational ensembles. Ensemble comparison and overlap were quantified by the Jensen-Shannon divergence and a new measure introduced here, the embedding depth, which determines the extent to which a given ensemble is subsumed by another ensemble and which may be a more useful measure in developing immunogens that confer conformational selectivity to an antibody.
Keywords: Cyclic peptides; ensemble similarity; epitope scaffolding; molecular dynamics; protein misfolding; virtual screening.
Conflict of interest statement
The authors declare the following competing financial interest(s): S.S.P. was Chief Physics Officer of ProMIS Neurosciences until October 2020. N.R.C. is Chief Scientific Officer of ProMIS Neurosciences. S.S.P., N.R.C., and X.P. are co-inventors on international patent application PCT/CA2019/051434 (Publication WO/2020/073121, applicant being University of British Columbia). The patent application describes immunogens and epitopes in alpha-synuclein, antibodies to these epitopes, and methods of their making as well as their use. Patent applications owned by the University of British Columbia are licensed to ProMIS Neurosciences. The work presented was financially supported in part by ProMIS Neurosciences. S.C.C.H., A.A., X.P., A.Yu.R., N.R.C., and S.S.P. have received consultation compensation from ProMIS Neurosciences.
Figures
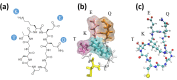



 and three
other scaffold properties: dynamic
flexibility (RMSF) of an epitope, total residue length of the cyclic
peptide scaffold, and the ranking, for α-synuclein epitopes
(a) EKTK, (b) KTKE, and (c) TKEQ. The number in each square is the
Pearson correlation coefficient.
and three
other scaffold properties: dynamic
flexibility (RMSF) of an epitope, total residue length of the cyclic
peptide scaffold, and the ranking, for α-synuclein epitopes
(a) EKTK, (b) KTKE, and (c) TKEQ. The number in each square is the
Pearson correlation coefficient.
 , and (d) rank and
, and (d) rank and  , for α-synuclein
epitope scaffolds.
The Pearson correlation coefficient r and the corresponding p-values are given for EKTK (green triangles), KTKE (purple
circles), and TKEQ (blue stars) scaffolds. The shaded areas around
the fitted lines are the 68% confidence intervals corresponding to
the standard errors.
, for α-synuclein
epitope scaffolds.
The Pearson correlation coefficient r and the corresponding p-values are given for EKTK (green triangles), KTKE (purple
circles), and TKEQ (blue stars) scaffolds. The shaded areas around
the fitted lines are the 68% confidence intervals corresponding to
the standard errors.
 and SASA exposure (f(SASAoff-target-exceed-cyclic) > 5%).
and SASA exposure (f(SASAoff-target-exceed-cyclic) > 5%).

References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Medical

