Identification of Key Pyroptosis-Related Genes and Distinct Pyroptosis-Related Clusters in Periodontitis
- PMID: 35844512
- PMCID: PMC9281553
- DOI: 10.3389/fimmu.2022.862049
Identification of Key Pyroptosis-Related Genes and Distinct Pyroptosis-Related Clusters in Periodontitis
Abstract
Aim: This study aims to identify pyroptosis-related genes (PRGs), their functional immune characteristics, and distinct pyroptosis-related clusters in periodontitis.
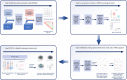
Methods: Differentially expressed (DE)-PRGs were determined by merging the expression profiles of GSE10334, GSE16134, and PRGs obtained from previous literatures and Molecular Signatures Database (MSigDB). Least absolute shrinkage and selection operator (LASSO) regression was applied to screen the prognostic PRGs and develop a prognostic model. Consensus clustering was applied to determine the pyroptosis-related clusters. Functional analysis and single-sample gene set enrichment analysis (ssGSEA) were performed to explore the biological characteristics and immune activities of the clusters. The hub pyroptosis-related modules were defined using weighted correlation network analysis (WGCNA).
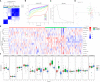
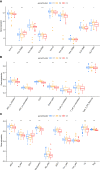
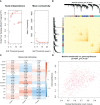
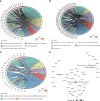
Results: Of the 26 periodontitis-related DE-PRGs, the highest positive relevance was for High-Mobility Group Box 1 (HMGB1) and SR-Related CTD Associated Factor 11 (SCAF11). A 14-PRG-based signature was developed through the LASSO model. In addition, three pyroptosis-related clusters were obtained based on the 14 prognostic PRGs. Caspase 3 (CASP3), Granzyme B (GZMB), Interleukin 1 Alpha (IL1A), IL1Beta (B), IL6, Phospholipase C Gamma 1 (PLCG1) and PYD And CARD Domain Containing (PYCARD) were dysregulated in the three clusters. Distinct biological functions and immune activities, including human leukocyte antigen (HLA) gene expression, immune cell infiltration, and immune pathway activities, were identified in the three pyroptosis-related clusters of periodontitis. Furthermore, the pink module associated with endoplasmic stress-related functions was found to be correlated with cluster 2 and was suggested as the hub pyroptosis-related module.
Conclusion: The study identified 14 key pyroptosis-related genes, three distinct pyroptosis-related clusters, and one pyroptosis-related gene module describing several molecular aspects of pyroptosis in the pathogenesis and immune micro-environment regulation of periodontitis and also highlighted functional heterogeneity in pyroptosis-related mechanisms.
Keywords: immune microenvironment; inflammasome; periodontitis; prognostic; pyroptosis.
Copyright © 2022 Ning, Acharya, Li, Schmalz and Huang.
Conflict of interest statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Figures



Similar articles
-
Relevance of pyroptosis-associated genes in nasopharyngeal carcinoma diagnosis and subtype classification.J Gene Med. 2024 Jan;26(1):e3653. doi: 10.1002/jgm.3653. J Gene Med. 2024. PMID: 38282154
-
Roles of Pyroptosis-Related Genes in the Diagnosis and Subtype Classification of Periodontitis.J Immunol Res. 2023 Apr 10;2023:8757233. doi: 10.1155/2023/8757233. eCollection 2023. J Immunol Res. 2023. PMID: 37090158 Free PMC article.
-
Exploring hub pyroptosis-related genes, molecular subtypes, and potential drugs in ankylosing spondylitis by comprehensive bioinformatics analysis and molecular docking.BMC Musculoskelet Disord. 2023 Jun 29;24(1):532. doi: 10.1186/s12891-023-06664-8. BMC Musculoskelet Disord. 2023. PMID: 37386410 Free PMC article.
-
Deciphering the role of pyroptosis-related genes and natural killer T cells in sepsis pathogenesis: a comprehensive bioinformatics and Mendelian randomization analysis.J Physiol Pharmacol. 2025 Apr;76(2). doi: 10.26402/jpp.2025.2.10. Epub 2025 May 5. J Physiol Pharmacol. 2025. PMID: 40350654
-
Pyroptosis-Related Gene Signature Predicts Prognosis and Indicates Immune Microenvironment Infiltration in Glioma.Front Cell Dev Biol. 2022 Apr 25;10:862493. doi: 10.3389/fcell.2022.862493. eCollection 2022. Front Cell Dev Biol. 2022. PMID: 35547808 Free PMC article.
Cited by
-
GLS as a Key Cuproptosis-Related Gene in Periodontitis: Insights from Single-Cell RNA Sequencing.Int Dent J. 2025 Apr;75(2):1360-1369. doi: 10.1016/j.identj.2024.10.001. Epub 2024 Oct 20. Int Dent J. 2025. PMID: 39428264 Free PMC article.
-
Estimation of noncanonical pyroptosis biomarkers gasdermin D and caspase 4 in gingiva of periodontitis and diabetes patients: An observational cross-sectional study.J Indian Soc Periodontol. 2024 May-Jun;28(3):297-303. doi: 10.4103/jisp.jisp_92_24. Epub 2024 Dec 2. J Indian Soc Periodontol. 2024. PMID: 39742067 Free PMC article.
-
ToxDAR: A Workflow Software for Analyzing Toxicologically Relevant Proteomic and Transcriptomic Data, from Data Preparation to Toxicological Mechanism Elucidation.Int J Mol Sci. 2024 Sep 2;25(17):9544. doi: 10.3390/ijms25179544. Int J Mol Sci. 2024. PMID: 39273492 Free PMC article.
-
Identification and validation of a novel prognostic signature based on mitochondria and oxidative stress related genes for glioblastoma.J Transl Med. 2023 Feb 22;21(1):136. doi: 10.1186/s12967-023-03970-6. J Transl Med. 2023. PMID: 36814293 Free PMC article.
-
Identification of ferroptosis, necroptosis, and pyroptosis-associated genes in periodontitis-affected human periodontal tissue using integrated bioinformatic analysis.Front Pharmacol. 2023 Jan 6;13:1098851. doi: 10.3389/fphar.2022.1098851. eCollection 2022. Front Pharmacol. 2023. PMID: 36686646 Free PMC article.
References
Publication types
MeSH terms
LinkOut - more resources
Full Text Sources
Research Materials
Miscellaneous

