Role of Nuclear Lamin A/C in the Regulation of Nav1.5 Channel and Microtubules: Lesson From the Pathogenic Lamin A/C Variant Q517X
- PMID: 35846372
- PMCID: PMC9277463
- DOI: 10.3389/fcell.2022.918760
Role of Nuclear Lamin A/C in the Regulation of Nav1.5 Channel and Microtubules: Lesson From the Pathogenic Lamin A/C Variant Q517X
Abstract
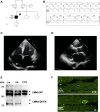
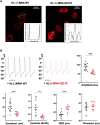
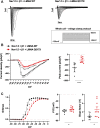
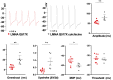
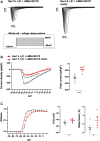
In this work, we studied an lmna nonsense mutation encoding for the C-terminally truncated Lamin A/C (LMNA) variant Q517X, which was described in patients affected by a severe arrhythmogenic cardiomyopathy with history of sudden death. We found that LMNA Q517X stably expressed in HL-1 cardiomyocytes abnormally aggregates at the nuclear envelope and within the nucleoplasm. Whole-cell patch clamp experiments showed that LMNA Q517X-expressing cardiomyocytes generated action potentials with reduced amplitude, overshoot, upstroke velocity and diastolic potential compared with LMNA WT-expressing cardiomyocytes. Moreover, the unique features of these cardiomyocytes were 1) hyper-polymerized tubulin network, 2) upregulated acetylated α-tubulin, and 3) cell surface Nav1.5 downregulation. These findings pointed the light on the role of tubulin and Nav1.5 channel in the abnormal electrical properties of LMNA Q517X-expressing cardiomyocytes. When expressed in HEK293 with Nav1.5 and its β1 subunit, LMNA Q517X reduced the peak Na+ current (INa) up to 63% with a shift toward positive potentials in the activation curve of the channel. Of note, both AP properties in cardiomyocytes and Nav1.5 kinetics in HEK293 cells were rescued in LMNA Q517X-expressing cells upon treatment with colchicine, an FDA-approved inhibitor of tubulin assembly. In conclusion, LMNA Q517X expression is associated with hyper-polymerization and hyper-acetylation of tubulin network with concomitant downregulation of Nav1.5 cell expression and activity, thus revealing 1) new mechanisms by which LMNA may regulate channels at the cell surface in cardiomyocytes and 2) new pathomechanisms and therapeutic targets in cardiac laminopathies.
Keywords: arrhythmias; electrophysiology; lamin A/C; sodium channel; tubulin.
Copyright © 2022 De Zio, Pietrafesa, Milano, Procino, Bramerio, Pepe, Forleo, Favale, Svelto, Gerbino and Carmosino.
Conflict of interest statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Figures



Similar articles
-
Functional Characterization of a Novel Truncating Mutation in Lamin A/C Gene in a Family with a Severe Cardiomyopathy with Conduction Defects.Cell Physiol Biochem. 2017;44(4):1559-1577. doi: 10.1159/000485651. Epub 2017 Dec 4. Cell Physiol Biochem. 2017. PMID: 29197877
-
Phosphorylation of Lamin A/C at serine 22 modulates Nav 1.5 function.Physiol Rep. 2021 Nov;9(22):e15121. doi: 10.14814/phy2.15121. Physiol Rep. 2021. PMID: 34806324 Free PMC article.
-
Clinical and functional characterization of a novel mutation in lamin a/c gene in a multigenerational family with arrhythmogenic cardiac laminopathy.PLoS One. 2015 Apr 2;10(4):e0121723. doi: 10.1371/journal.pone.0121723. eCollection 2015. PLoS One. 2015. PMID: 25837155 Free PMC article.
-
Dysfunctional Nav1.5 channels due to SCN5A mutations.Exp Biol Med (Maywood). 2018 Jun;243(10):852-863. doi: 10.1177/1535370218777972. Epub 2018 May 27. Exp Biol Med (Maywood). 2018. PMID: 29806494 Free PMC article. Review.
-
Lamin A/C Cardiomyopathies: Current Understanding and Novel Treatment Strategies.Curr Treat Options Cardiovasc Med. 2017 Mar;19(3):21. doi: 10.1007/s11936-017-0520-z. Curr Treat Options Cardiovasc Med. 2017. PMID: 28299614 Review.
Cited by
-
Targeting unfolded protein response reverts ER stress and ER Ca2+ homeostasis in cardiomyocytes expressing the pathogenic variant of Lamin A/C R321X.J Transl Med. 2023 May 22;21(1):340. doi: 10.1186/s12967-023-04170-y. J Transl Med. 2023. PMID: 37217929 Free PMC article.
References
LinkOut - more resources
Full Text Sources
Miscellaneous

