A Hypoxia-Related lncRNA Signature Correlates with Survival and Tumor Microenvironment in Colorectal Cancer
- PMID: 35846431
- PMCID: PMC9286950
- DOI: 10.1155/2022/9935705
A Hypoxia-Related lncRNA Signature Correlates with Survival and Tumor Microenvironment in Colorectal Cancer
Abstract
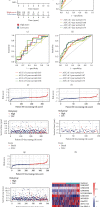
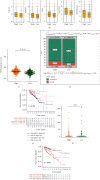
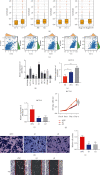
The hypoxic tumor microenvironment and long noncoding RNAs (lncRNAs) are pivotal in cancer progression and correlate with the survival outcome of patients. However, the role of hypoxia-related lncRNAs (HRLs) in colorectal cancer (CRC) development remains largely unknown. Herein, we developed a hypoxia-related lncRNA signature to predict patients' survival and immune infiltration. The RNA-sequencing data of 500 CRC patients were obtained from The Cancer Genome Atlas (TCGA) dataset, and HRLs were selected using Pearson's analysis. Next, the Cox regression analysis was applied to construct a risk signature consisting of 9 HRLs. This signature could predict the overall survival (OS) of CRC patients with high accuracy in training, validation, and entire cohort. This signature was an independent risk factor and exerted predictive ability in different subgroups. Functional analysis revealed different molecular features between high- and low-risk groups. A series of drugs including cisplatin showed different sensitivities between the two groups. The expression pattern of immune checkpoints was also distinct between the two clusters in this model. Furthermore, the high-risk group had higher immune, stromal, and ESTIMATE score and a more repressive immune microenvironment than the low-risk group. Moreover, MYOSLID, one of the lncRNAs in this signature, could significantly regulate the proliferation, invasion, and metastasis of CRC.
Copyright © 2022 Xinyang Zhong et al.
Conflict of interest statement
The authors declare that they have no competing interests.
Figures





References
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Medical

