Frequent convergence of mcr-9 and carbapenemase genes in Enterobacter cloacae complex driven by epidemic plasmids and host incompatibility
- PMID: 35848148
- PMCID: PMC9359198
- DOI: 10.1080/22221751.2022.2103456
Frequent convergence of mcr-9 and carbapenemase genes in Enterobacter cloacae complex driven by epidemic plasmids and host incompatibility
Abstract
Convergence of mcr and carbapenemase genes has been sporadically detected in Enterobacter cloacae complex (ECC) with an upward trend. However, the state of the epidemic and underlying mechanism of such convergence has been poorly understood. In this study, the co-occurrence of MCR and carbapenemases was systematically analyzed in 230 clinical ECC isolates collected between 2000 and 2018 together with a global dataset consisting of 3,559 ECC genomes compiled from GenBank. We identified 48 mcr-9/mcr-10-positive isolates (MCR-ECC) (20.9%) in our collection, and a comparable ratio of MCR-ECC (720/3559, 20.2%) was detected in the global dataset. A high prevalence of carbapenemase-producing MCR-ECC (MCR-CREC) was further identified in the MCR-ECC of both datasets (16/48, 33.3%; 388/720, 53.9%), demonstrating a frequent convergence of mcr-9/10 and carbapenemase genes in ECC worldwide. An epidemic IncHI2/2A plasmid with a highly conserved backbone was identified and largely contributed to the dissemination of mcr-9 in ECC worldwide. A highly conserved IncX3-type NDM-1-carrying plasmid and IncN-type IMP-4-carrying plasmid were additionally detected in MCR-CREC isolated in China. Our surveillance data showed that MCR-CREC emerged (in 2013) much later than MCR-ECC (in 2000), indicating that MCR-CREC could be derived from MCR-ECC by additional captures of carbapenemase-encoding plasmids. Tests of plasmid stability and incompatibility showed that the mcr-9/mcr-10-encoding plasmids with the NDM-1-encoding plasmids stably remained in ECC but incompatible in Escherichia coli, suggesting that the convergence was host-dependent. The findings extend our concern on the convergence of resistance to the last resort antibiotics and highlight the necessity of continued surveillance in the future.
Keywords: Enterobacter cloacae complex; antibiotic resistance; carbapenemase; epidemic plasmid; mcr-9.
Conflict of interest statement
No potential conflict of interest was reported by the author(s).
Figures

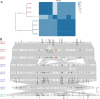
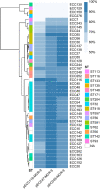
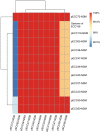

Similar articles
-
Emergence of concurrently transmissible mcr-9 and carbapenemase genes in bloodborne colistin-resistant Enterobacter cloacae complex isolated from ICU patients in Kolkata, India.Microbiol Spectr. 2025 Mar 4;13(3):e0154224. doi: 10.1128/spectrum.01542-24. Epub 2025 Feb 6. Microbiol Spectr. 2025. PMID: 39912656 Free PMC article.
-
Molecular characterization of NDM-1-producing carbapenem-resistant E. cloacae complex from a tertiary hospital in Chongqing, China.Front Cell Infect Microbiol. 2022 Aug 8;12:935165. doi: 10.3389/fcimb.2022.935165. eCollection 2022. Front Cell Infect Microbiol. 2022. PMID: 36004335 Free PMC article.
-
Whole genome sequence-based molecular characterization of blood isolates of carbapenem-resistant Enterobacter cloacae complex from ICU patients in Kolkata, India, during 2017-2022: emergence of phylogenetically heterogeneous Enterobacter hormaechei subsp. xiangfangensis.Microbiol Spectr. 2024 Apr 2;12(4):e0352923. doi: 10.1128/spectrum.03529-23. Epub 2024 Feb 22. Microbiol Spectr. 2024. PMID: 38385742 Free PMC article.
-
Global prevalence, characteristics, and future prospects of IncX3 plasmids: A review.Front Microbiol. 2022 Sep 6;13:979558. doi: 10.3389/fmicb.2022.979558. eCollection 2022. Front Microbiol. 2022. PMID: 36147856 Free PMC article. Review.
-
Carbapenem Resistance in Gram-Negative Bacteria: The Not-So-Little Problem in the Little Red Dot.Microorganisms. 2016 Feb 16;4(1):13. doi: 10.3390/microorganisms4010013. Microorganisms. 2016. PMID: 27681907 Free PMC article. Review.
Cited by
-
High prevalence of colistin heteroresistance in specific species and lineages of Enterobacter cloacae complex derived from human clinical specimens.Ann Clin Microbiol Antimicrob. 2023 Jul 15;22(1):60. doi: 10.1186/s12941-023-00610-1. Ann Clin Microbiol Antimicrob. 2023. PMID: 37454128 Free PMC article.
-
Novel plasmid-mediated CMY variant (CMY-192) conferring ceftazidime-avibactam resistance in multidrug-resistant Escherichia coli.Antimicrob Agents Chemother. 2024 Dec 5;68(12):e0090624. doi: 10.1128/aac.00906-24. Epub 2024 Oct 29. Antimicrob Agents Chemother. 2024. PMID: 39470201 Free PMC article.
-
Molecular epidemiology and genomic characterization of a plasmid-mediated mcr-10 and blaNDM-1 co-harboring multidrug-resistant Enterobacter asburiae.Comput Struct Biotechnol J. 2023 Aug 6;21:3885-3893. doi: 10.1016/j.csbj.2023.08.004. eCollection 2023. Comput Struct Biotechnol J. 2023. PMID: 37602227 Free PMC article.
-
Insertion sequences in mgrB and mutations in two-component system genes confer high polymyxin resistance to carbapenem-resistant Enterobacter cloacae complex strains.Front Microbiol. 2025 Mar 17;16:1553148. doi: 10.3389/fmicb.2025.1553148. eCollection 2025. Front Microbiol. 2025. PMID: 40165791 Free PMC article.
-
Whole-Genome Analysis of blaNDM-Bearing Proteus mirabilis Isolates and mcr-1-Positive Escherichia coli Isolates Carrying blaNDM from the Same Fresh Vegetables in China.Foods. 2023 Jan 20;12(3):492. doi: 10.3390/foods12030492. Foods. 2023. PMID: 36766021 Free PMC article.
References
-
- Giamarellou H, Poulakou G.. Multidrug-Resistant gram-negative infections. Drugs. 2009;69:1879–1901. - PubMed
-
- D'Onofrio V, Conzemius R, Varda-Brkic D, et al. . Epidemiology of colistin-resistant, carbapenemase-producing enterobacteriaceae and acinetobacter baumannii in Croatia. Infect Genet Evol. 2020;81:104263. - PubMed
-
- Kieffer N, Ahmed MO, Elramalli AK, et al. . Colistin-resistant carbapenemase-producing isolates among klebsiella spp. and acinetobacter baumannii in Tripoli, Libya. J Glob Antimicrob Resist. 2018;13:37–39. - PubMed
-
- Liu Y-Y, Wang Y, Walsh TR, et al. . Emergence of plasmid-mediated colistin resistance mechanism MCR-1 in animals and human beings in China: a microbiological and molecular biological study. Lancet Infect Dis. 2016;16:161–168. - PubMed
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
