Genomic and epigenetic landscapes drive CRISPR-based genome editing in Bifidobacterium
- PMID: 35857876
- PMCID: PMC9335239
- DOI: 10.1073/pnas.2205068119
Genomic and epigenetic landscapes drive CRISPR-based genome editing in Bifidobacterium
Abstract
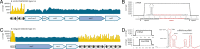
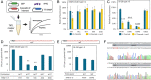
Bifidobacterium is a commensal bacterial genus ubiquitous in the human gastrointestinal tract, which is associated with a range of health benefits. The advent of CRISPR-based genome editing technologies provides opportunities to investigate the genetics of important bacteria and transcend the lack of genetic tools in bifidobacteria to study the basis for their health-promoting attributes. Here, we repurpose the endogenous type I-G CRISPR-Cas system and adopt an exogenous CRISPR base editor for genome engineering in B. animalis subsp. lactis, demonstrating that both genomic and epigenetic contexts drive editing outcomes across strains. We reprogrammed the endogenous type I-G system to screen for naturally occurring large deletions up to 27 kb and to generate a 500-bp deletion in tetW to abolish tetracycline resistance. A CRISPR-cytosine base editor was optimized to install C•G-to-T•A amber mutations to resensitize multiple B. lactis strains to tetracycline. Remarkably, we uncovered epigenetic patterns that are distributed unevenly among B. lactis strains, despite their genomic homogeneity, that may contribute to editing efficiency variability. Insights were also expanded to Bifidobacterium longum subsp. infantis to emphasize the broad relevance of these findings. This study highlights the need to develop individualized CRISPR-based genome engineering approaches for distinct bacterial strains and opens avenues for engineering of next generation probiotics.
Keywords: CRISPR-Cas; bifidobacterium; epigenomics; genomics; probiotics.
Conflict of interest statement
Competing interest statement: R.B., C.H.-C. and Y.J.G are inventors on several patents related to CRISPR-Cas systems and their uses. R.B. is a cofounder of Intellia Therapeutics, Locus Biosciences, TreeCo, CRISPR Biotechnologies and Ancilia Biosciences, and a shareholder of Caribou Biosciences, Inari Ag, Felix Biotechnologies, and Provaxus. C.H.-C. is a cofounder of Microviable Therapeutics and shareholder of CRISPR Biotechnologies. W.M. is employed by IFF Health & Biosciences, International Flavors and Fragrances, Inc., which commercializes probiotic products. R.B. and J.v.d.O. are co-authors on a 2018 news and views article and a 2020 review article.
Figures




Similar articles
-
Highly efficient CRISPR-Cas9 base editing in Bifidobacterium with bypass of restriction modification systems.Appl Environ Microbiol. 2025 Apr 23;91(4):e0198524. doi: 10.1128/aem.01985-24. Epub 2025 Mar 10. Appl Environ Microbiol. 2025. PMID: 40062897 Free PMC article.
-
Harnessing the endogenous Type I-C CRISPR-Cas system for genome editing in Bifidobacterium breve.Appl Environ Microbiol. 2024 Mar 20;90(3):e0207423. doi: 10.1128/aem.02074-23. Epub 2024 Feb 6. Appl Environ Microbiol. 2024. PMID: 38319094 Free PMC article.
-
CRISPR-based engineering of next-generation lactic acid bacteria.Curr Opin Microbiol. 2017 Jun;37:79-87. doi: 10.1016/j.mib.2017.05.015. Epub 2017 Jun 13. Curr Opin Microbiol. 2017. PMID: 28622636 Review.
-
Portable CRISPR-Cas9N System for Flexible Genome Engineering in Lactobacillus acidophilus, Lactobacillus gasseri, and Lactobacillus paracasei.Appl Environ Microbiol. 2021 Feb 26;87(6):e02669-20. doi: 10.1128/AEM.02669-20. Print 2021 Feb 26. Appl Environ Microbiol. 2021. PMID: 33397707 Free PMC article.
-
Advancing mechanistic understanding and bioengineering of probiotic lactobacilli and bifidobacteria by genome editing.Curr Opin Biotechnol. 2021 Aug;70:75-82. doi: 10.1016/j.copbio.2020.12.015. Epub 2021 Jan 11. Curr Opin Biotechnol. 2021. PMID: 33445135 Review.
Cited by
-
A highly efficient method for genomic deletion across diverse lengths in thermophilic Parageobacillus thermoglucosidasius.Synth Syst Biotechnol. 2024 May 17;9(4):658-666. doi: 10.1016/j.synbio.2024.05.009. eCollection 2024 Dec. Synth Syst Biotechnol. 2024. PMID: 38817825 Free PMC article.
-
Type I CRISPR-Cas-mediated microbial gene editing and regulation.AIMS Microbiol. 2023 Dec 18;9(4):780-800. doi: 10.3934/microbiol.2023040. eCollection 2023. AIMS Microbiol. 2023. PMID: 38173969 Free PMC article. Review.
-
The therapeutic value of bifidobacteria in cardiovascular disease.NPJ Biofilms Microbiomes. 2023 Oct 30;9(1):82. doi: 10.1038/s41522-023-00448-7. NPJ Biofilms Microbiomes. 2023. PMID: 37903770 Free PMC article. Review.
-
CRISPR/FnCas12a-mediated efficient multiplex and iterative genome editing in bacterial plant pathogens without donor DNA templates.PLoS Pathog. 2023 Jan 10;19(1):e1010961. doi: 10.1371/journal.ppat.1010961. eCollection 2023 Jan. PLoS Pathog. 2023. PMID: 36626407 Free PMC article.
-
In situ targeted base editing of bacteria in the mouse gut.Nature. 2024 Aug;632(8026):877-884. doi: 10.1038/s41586-024-07681-w. Epub 2024 Jul 10. Nature. 2024. PMID: 38987595 Free PMC article.
References
-
- Fukuda S., et al. , Bifidobacteria can protect from enteropathogenic infection through production of acetate. Nature 469, 543–547 (2011). - PubMed
-
- O’Mahony L., et al. , Lactobacillus and bifidobacterium in irritable bowel syndrome: symptom responses and relationship to cytokine profiles. Gastroenterology 128, 541–551 (2005). - PubMed
-
- Henrick B. M., et al. , Bifidobacteria-mediated immune system imprinting early in life. Cell 184, 3884–3898.e11 (2021). - PubMed
Publication types
MeSH terms
LinkOut - more resources
Full Text Sources

