Phase Separation and Correlated Motions in Motorized Genome
- PMID: 35858189
- PMCID: PMC9899348
- DOI: 10.1021/acs.jpcb.2c03238
Phase Separation and Correlated Motions in Motorized Genome
Abstract
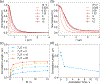
The human genome is arranged in the cell nucleus nonrandomly, and phase separation has been proposed as an important driving force for genome organization. However, the cell nucleus is an active system, and the contribution of nonequilibrium activities to phase separation and genome structure and dynamics remains to be explored. We simulated the genome using an energy function parametrized with chromosome conformation capture (Hi-C) data with the presence of active, nondirectional forces that break the detailed balance. We found that active forces that may arise from transcription and chromatin remodeling can dramatically impact the spatial localization of heterochromatin. When applied to euchromatin, active forces can drive heterochromatin to the nuclear envelope and compete with passive interactions among heterochromatin that tend to pull them in opposite directions. Furthermore, active forces induce long-range spatial correlations among genomic loci beyond single chromosome territories. We further showed that the impact of active forces could be understood from the effective temperature defined as the fluctuation-dissipation ratio. Our study suggests that nonequilibrium activities can significantly impact genome structure and dynamics, producing unexpected collective phenomena.
Figures
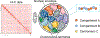



Similar articles
-
Heterochromatin drives compartmentalization of inverted and conventional nuclei.Nature. 2019 Jun;570(7761):395-399. doi: 10.1038/s41586-019-1275-3. Epub 2019 Jun 5. Nature. 2019. PMID: 31168090 Free PMC article.
-
The interplay of chromatin phase separation and lamina interactions in nuclear organization.Biophys J. 2021 Nov 16;120(22):5005-5017. doi: 10.1016/j.bpj.2021.10.012. Epub 2021 Oct 13. Biophys J. 2021. PMID: 34653387 Free PMC article.
-
How to rule the nucleus: divide et impera.Curr Opin Cell Biol. 2016 Jun;40:47-59. doi: 10.1016/j.ceb.2016.02.014. Epub 2016 Mar 1. Curr Opin Cell Biol. 2016. PMID: 26938331 Review.
-
The genome organization of Neurospora crassa at high resolution uncovers principles of fungal chromosome topology.G3 (Bethesda). 2022 May 6;12(5):jkac053. doi: 10.1093/g3journal/jkac053. G3 (Bethesda). 2022. PMID: 35244156 Free PMC article.
-
Nuclear dynamics of radiation-induced foci in euchromatin and heterochromatin.Mutat Res. 2013 Oct;750(1-2):56-66. doi: 10.1016/j.mrfmmm.2013.08.001. Epub 2013 Aug 16. Mutat Res. 2013. PMID: 23958412 Free PMC article. Review.
Cited by
-
Large-scale data-driven and physics-based models offer insights into the relationships among the structures, dynamics, and functions of chromosomes.J Mol Cell Biol. 2023 Nov 27;15(6):mjad042. doi: 10.1093/jmcb/mjad042. J Mol Cell Biol. 2023. PMID: 37365687 Free PMC article. Review.
-
Transcription-induced active forces suppress chromatin motion.Proc Natl Acad Sci U S A. 2024 Mar 19;121(12):e2307309121. doi: 10.1073/pnas.2307309121. Epub 2024 Mar 15. Proc Natl Acad Sci U S A. 2024. PMID: 38489381 Free PMC article.
-
Polymer folding through active processes recreates features of genome organization.Proc Natl Acad Sci U S A. 2023 May 16;120(20):e2221726120. doi: 10.1073/pnas.2221726120. Epub 2023 May 8. Proc Natl Acad Sci U S A. 2023. PMID: 37155885 Free PMC article.
-
Molecular Determinants for the Layering and Coarsening of Biological Condensates.Aggregate (Hoboken). 2022 Dec;3(6):e306. doi: 10.1002/agt2.306. Epub 2022 Dec 10. Aggregate (Hoboken). 2022. PMID: 37065433 Free PMC article.
-
Explicit Ion Modeling Predicts Physicochemical Interactions for Chromatin Organization.bioRxiv [Preprint]. 2023 Nov 15:2023.05.16.541030. doi: 10.1101/2023.05.16.541030. bioRxiv. 2023. Update in: Elife. 2024 Jan 30;12:RP90073. doi: 10.7554/eLife.90073. PMID: 37293007 Free PMC article. Updated. Preprint.
References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

