Single-molecule analysis of actin filament debranching by cofilin and GMF
- PMID: 35858314
- PMCID: PMC9304009
- DOI: 10.1073/pnas.2115129119
Single-molecule analysis of actin filament debranching by cofilin and GMF
Abstract
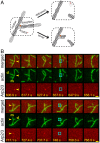
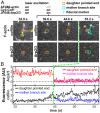
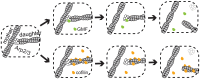
Eukaryotic cells contain branched actin networks that are essential for endocytosis, motility, and other key cellular processes. These networks, which are formed by filamentous actin and the Arp2/3 complex, must subsequently be debranched to allow network remodeling and to recycle the Arp2/3 complex. Debranching appears to be catalyzed by two different members of the actin depolymerizing factor homology protein family: cofilin and glial maturation factor (GMF). However, their mechanisms of debranching are only partially understood. Here, we used single-molecule fluorescence imaging of Arp2/3 complex and actin filaments under physiological ionic conditions to observe debranching by GMF and cofilin. We demonstrate that cofilin, like GMF, is an authentic debrancher independent of its filament-severing activity and that the debranching activities of the two proteins are additive. While GMF binds directly to the Arp2/3 complex, cofilin selectively accumulates on branch-junction daughter filaments in tropomyosin-decorated networks just prior to debranching events. Quantitative comparison of debranching rates with the known kinetics of cofilin-actin binding suggests that cofilin occupancy of a particular single actin site at the branch junction is sufficient to trigger debranching. In rare cases in which the order of departure could be resolved during GMF- or cofilin-induced debranching, the Arp2/3 complex left the branch junction bound to the pointed end of the daughter filament, suggesting that both GMF and cofilin can work by destabilizing the mother filament-Arp2/3 complex interface. Taken together, these observations suggest that GMF and cofilin promote debranching by distinct yet complementary mechanisms.
Keywords: Arp2/3 complex; actin depolymerizing factor homology; branched actin networks; glial maturation factor; tropomyosin.
Conflict of interest statement
The authors declare no competing interest.
Figures



Similar articles
-
GMF is a cofilin homolog that binds Arp2/3 complex to stimulate filament debranching and inhibit actin nucleation.Curr Biol. 2010 May 11;20(9):861-7. doi: 10.1016/j.cub.2010.03.026. Epub 2010 Apr 1. Curr Biol. 2010. PMID: 20362448 Free PMC article.
-
GMF severs actin-Arp2/3 complex branch junctions by a cofilin-like mechanism.Curr Biol. 2013 Jun 17;23(12):1037-45. doi: 10.1016/j.cub.2013.04.058. Epub 2013 May 30. Curr Biol. 2013. PMID: 23727094 Free PMC article.
-
Glia maturation factor (GMF) interacts with Arp2/3 complex in a nucleotide state-dependent manner.J Biol Chem. 2013 Sep 6;288(36):25683-25688. doi: 10.1074/jbc.C113.493338. Epub 2013 Jul 29. J Biol Chem. 2013. PMID: 23897816 Free PMC article.
-
GMF as an Actin Network Remodeling Factor.Trends Cell Biol. 2018 Sep;28(9):749-760. doi: 10.1016/j.tcb.2018.04.008. Epub 2018 May 18. Trends Cell Biol. 2018. PMID: 29779865 Free PMC article. Review.
-
Biophysics of actin filament severing by cofilin.FEBS Lett. 2013 Apr 17;587(8):1215-9. doi: 10.1016/j.febslet.2013.01.062. Epub 2013 Feb 5. FEBS Lett. 2013. PMID: 23395798 Free PMC article. Review.
Cited by
-
Mechanisms of actin disassembly and turnover.J Cell Biol. 2023 Dec 4;222(12):e202309021. doi: 10.1083/jcb.202309021. Epub 2023 Nov 10. J Cell Biol. 2023. PMID: 37948068 Free PMC article. Review.
-
Cortactin stabilizes actin branches by bridging activated Arp2/3 to its nucleated actin filament.Nat Struct Mol Biol. 2024 May;31(5):801-809. doi: 10.1038/s41594-023-01205-2. Epub 2024 Jan 24. Nat Struct Mol Biol. 2024. PMID: 38267598 Free PMC article.
-
The stabilization of Arp2/3 complex generated actin filaments.Biochem Soc Trans. 2024 Feb 28;52(1):343-352. doi: 10.1042/BST20230638. Biochem Soc Trans. 2024. PMID: 38288872 Free PMC article. Review.
-
A natural hydrogel complex improves intervertebral disc degeneration by correcting fatty acid metabolism and inhibiting nucleus pulposus cell pyroptosis.Mater Today Bio. 2024 May 3;26:101081. doi: 10.1016/j.mtbio.2024.101081. eCollection 2024 Jun. Mater Today Bio. 2024. PMID: 38741924 Free PMC article.
-
Regeneration of actin filament branches from the same Arp2/3 complex.Sci Adv. 2024 Jan 26;10(4):eadj7681. doi: 10.1126/sciadv.adj7681. Epub 2024 Jan 26. Sci Adv. 2024. PMID: 38277459 Free PMC article.
References
-
- Senju Y., Lappalainen P., Regulation of actin dynamics by PI(4,5)P2 in cell migration and endocytosis. Curr. Opin. Cell Biol. 56, 7–13 (2019). - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

