Single-cell entropy network detects the activity of immune cells based on ribosomal protein genes
- PMID: 35860411
- PMCID: PMC9287362
- DOI: 10.1016/j.csbj.2022.06.056
Single-cell entropy network detects the activity of immune cells based on ribosomal protein genes
Abstract
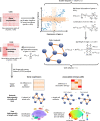
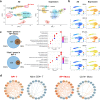
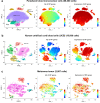
We developed a new computational method, Single-Cell Entropy Network (SCEN) to analyze single-cell RNA-seq data, which used the information of gene-gene associations to discover new heterogeneity of immune cells as well as identify existing cell types. Based on SCEN, we defined association-entropy (AE) for each cell and each gene through single-cell gene co-expression networks to measure the strength of association between each gene and all other genes at a single-cell resolution. Analyses of public datasets indicated that the AE of ribosomal protein genes (RP genes) varied greatly even in the same cell type of immune cells and the average AE of RP genes of immune cells in each person was significantly associated with the healthy/disease state of this person. Based on existing research and theory, we inferred that the AE of RP genes represented the heterogeneity of ribosomes and reflected the activity of immune cells. We believe SCEN can provide more biological insights into the heterogeneity and diversity of immune cells, especially the change of immune cells in the diseases.
Keywords: Association-entropy; Immune cell; Ribosomal protein gene; Single-cell RNA-seq; Single-cell entropy network.
© 2022 The Author(s).
Conflict of interest statement
The authors declare that they have no known competing financial interests or personal relationships that could have appeared to influence the work reported in this paper.
Figures



Similar articles
-
SGAE: single-cell gene association entropy for revealing critical states of cell transitions during embryonic development.Brief Bioinform. 2023 Sep 22;24(6):bbad366. doi: 10.1093/bib/bbad366. Brief Bioinform. 2023. PMID: 37833841
-
MICRAT: a novel algorithm for inferring gene regulatory networks using time series gene expression data.BMC Syst Biol. 2018 Dec 14;12(Suppl 7):115. doi: 10.1186/s12918-018-0635-1. BMC Syst Biol. 2018. PMID: 30547796 Free PMC article.
-
[Quantifying the state of cell differentiation based on the gene networks entropy].Sheng Wu Gong Cheng Xue Bao. 2022 Feb 25;38(2):820-830. doi: 10.13345/j.cjb.210140. Sheng Wu Gong Cheng Xue Bao. 2022. PMID: 35234401 Chinese.
-
Single-cell network biology for resolving cellular heterogeneity in human diseases.Exp Mol Med. 2020 Nov;52(11):1798-1808. doi: 10.1038/s12276-020-00528-0. Epub 2020 Nov 26. Exp Mol Med. 2020. PMID: 33244151 Free PMC article. Review.
-
Ribosomal heterogeneity - A new inroad for pharmacological innovation.Biochem Pharmacol. 2020 May;175:113874. doi: 10.1016/j.bcp.2020.113874. Epub 2020 Feb 24. Biochem Pharmacol. 2020. PMID: 32105657 Review.
Cited by
-
Early life adversity is associated with differential gene expression in immune cells: A cluster-based analysis across an acute psychosocial stressor.Brain Behav Immun. 2024 Jul;119:724-733. doi: 10.1016/j.bbi.2024.04.035. Epub 2024 Apr 23. Brain Behav Immun. 2024. PMID: 38663776 Free PMC article.
-
A single-cell transcriptional landscape of immune cells shows disease-specific changes of T cell and macrophage populations in human achalasia.Nat Commun. 2023 Aug 4;14(1):4685. doi: 10.1038/s41467-023-39750-5. Nat Commun. 2023. PMID: 37542039 Free PMC article.
-
Direct microglia replacement reveals pathologic and therapeutic contributions of brain macrophages to a monogenic neurological disease.Immunity. 2025 May 13;58(5):1254-1268.e9. doi: 10.1016/j.immuni.2025.03.019. Epub 2025 Apr 30. Immunity. 2025. PMID: 40311614
-
The network structural entropy for single-cell RNA sequencing data during skin aging.Brief Bioinform. 2024 Nov 22;26(1):bbae698. doi: 10.1093/bib/bbae698. Brief Bioinform. 2024. PMID: 39757115 Free PMC article.
References
-
- Grün D., Lyubimova A., Kester L., Wiebrands K., Basak O., Sasaki N., et al. Single-cell messenger RNA sequencing reveals rare intestinal cell types. Nature. 2015;525(7568):251–255. - PubMed
-
- Shapiro E., Biezuner T., Linnarsson S. Single-cell sequencing-based technologies will revolutionize whole-organism science. Nat Rev Genet. 2013;14(9):618–630. - PubMed
Associated data
LinkOut - more resources
Full Text Sources
