Computational Modeling of Macrophage Iron Sequestration during Host Defense against Aspergillus
- PMID: 35862797
- PMCID: PMC9429928
- DOI: 10.1128/msphere.00074-22
Computational Modeling of Macrophage Iron Sequestration during Host Defense against Aspergillus
Abstract
Iron is essential to the virulence of Aspergillus species, and restricting iron availability is a critical mechanism of antimicrobial host defense. Macrophages recruited to the site of infection are at the crux of this process, employing multiple intersecting mechanisms to orchestrate iron sequestration from pathogens. To gain an integrated understanding of how this is achieved in aspergillosis, we generated a transcriptomic time series of the response of human monocyte-derived macrophages to Aspergillus and used this and the available literature to construct a mechanistic computational model of iron handling of macrophages during this infection. We found an overwhelming macrophage response beginning 2 to 4 h after exposure to the fungus, which included upregulated transcription of iron import proteins transferrin receptor-1, divalent metal transporter-1, and ZIP family transporters, and downregulated transcription of the iron exporter ferroportin. The computational model, based on a discrete dynamical systems framework, consisted of 21 3-state nodes, and was validated with additional experimental data that were not used in model generation. The model accurately captures the steady state and the trajectories of most of the quantitatively measured nodes. In the experimental data, we surprisingly found that transferrin receptor-1 upregulation preceded the induction of inflammatory cytokines, a feature that deviated from model predictions. Model simulations suggested that direct induction of transferrin receptor-1 (TfR1) after fungal recognition, independent of the iron regulatory protein-labile iron pool (IRP-LIP) system, explains this finding. We anticipate that this model will contribute to a quantitative understanding of iron regulation as a fundamental host defense mechanism during aspergillosis. IMPORTANCE Invasive pulmonary aspergillosis is a major cause of death among immunosuppressed individuals despite the best available therapy. Depriving the pathogen of iron is an essential component of host defense in this infection, but the mechanisms by which the host achieves this are complex. To understand how recruited macrophages mediate iron deprivation during the infection, we developed and validated a mechanistic computational model that integrates the available information in the field. The insights provided by this approach can help in designing iron modulation therapies as anti-fungal treatments.
Keywords: Aspergillus fumigatus; iron regulation; macrophage; mathematical model.
Conflict of interest statement
The authors declare no conflict of interest.
Figures


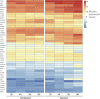



References
-
- Kontoyiannis DP, Marr KA, Park BJ, Alexander BD, Anaissie EJ, Walsh TJ, Ito J, Andes DR, Baddley JW, Brown JM, Brumble LM, Freifeld AG, Hadley S, Herwaldt LA, Kauffman CA, Knapp K, Lyon GM, Morrison VA, Papanicolaou G, Patterson TF, Perl TM, Schuster MG, Walker R, Wannemuehler KA, Wingard JR, Chiller TM, Pappas PG. 2010. Prospective surveillance for invasive fungal infections in hematopoietic stem cell transplant recipients, 2001–2006: overview of the Transplant-Associated Infection Surveillance Network (TRANSNET) Database. Clin Infect Dis 50:1091–1100. doi: 10.1086/651263. - DOI - PubMed
-
- Pappas PG, Alexander BD, Andes DR, Hadley S, Kauffman CA, Freifeld A, Anaissie EJ, Brumble LM, Herwaldt L, Ito J, Kontoyiannis DP, Lyon GM, Marr KA, Morrison VA, Park BJ, Patterson TF, Perl TM, Oster RA, Schuster MG, Walker R, Walsh TJ, Wannemuehler KA, Chiller TM. 2010. Invasive fungal infections among organ transplant recipients: results of the Transplant-Associated Infection Surveillance Network (TRANSNET). Clin Infect Dis 50:1101–1111. doi: 10.1086/651262. - DOI - PubMed
-
- Neofytos D, Horn D, Anaissie E, Steinbach W, Olyaei A, Fishman J, Pfaller M, Chang C, Webster K, Marr K. 2009. Epidemiology and outcome of invasive fungal infection in adult hematopoietic stem cell transplant recipients: analysis of Multicenter Prospective Antifungal Therapy (PATH) Alliance registry. Clin Infect Dis 48:265–273. doi: 10.1086/595846. - DOI - PubMed
-
- Neofytos D, Treadway S, Ostrander D, Alonso CD, Dierberg KL, Nussenblatt V, Durand CM, Thompson CB, Marr KA. 2013. Epidemiology, outcomes, and mortality predictors of invasive mold infections among transplant recipients: a 10-year, single-center experience. Transpl Infect Dis 15:233–242. doi: 10.1111/tid.12060. - DOI - PMC - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Medical
Molecular Biology Databases
