Genetic and Structural Variation in the O-Antigen of Salmonella enterica Serovar Typhimurium Isolates Causing Bloodstream Infections in the Democratic Republic of the Congo
- PMID: 35862803
- PMCID: PMC9426603
- DOI: 10.1128/mbio.00374-22
Genetic and Structural Variation in the O-Antigen of Salmonella enterica Serovar Typhimurium Isolates Causing Bloodstream Infections in the Democratic Republic of the Congo
Abstract
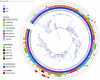
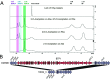
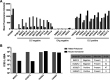
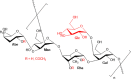
Salmonella enterica serovar Typhimurium causes a devastating burden of invasive disease in sub-Saharan Africa with high levels of antimicrobial resistance. No licensed vaccine is available, but O-antigen-based candidates are in development, as the O-antigen moiety of lipopolysaccharides is the principal target of protective immunity. The vaccines under development are designed based on isolates with O-antigen O-acetylated at position C-2 of abequose, giving the O:5 antigen. Serotyping data on recent Salmonella Typhimurium clinical isolates from the Democratic Republic of the Congo (DRC), however, indicate increasing levels of isolates without O:5. The importance and distribution of this loss of O:5 antigen in the population as well as the genetic mechanism responsible for the loss and chemical characteristics of the O-antigen are poorly understood. In this study, we Illumina whole-genome sequenced 354 Salmonella Typhimurium isolates from the DRC, which were isolated between 2002 and 2017. We used genomics and phylogenetics combined with chemical approaches (1H nuclear magnetic resonance [NMR], high-performance anion-exchange chromatography with pulsed amperometric detection [HPAEC-PAD], high-performance liquid chromatography-PAD [HPLC-PAD], and HPLC-size exclusion chromatography [HPLC-SEC]) to characterize the O-antigen features within the bacterial population. We observed convergent evolution toward the loss of the O:5 epitope predominantly caused by recombination events in a single gene, the O-acetyltransferase gene oafA. In addition, we observe further O-antigen variations, including O-acetylation of the rhamnose residue, different levels of glucosylation, and the absence of O-antigen repeating units. Large recombination events underlying O-antigen variation were resolved using long-read MinION sequencing. Our study suggests evolutionary pressure toward O-antigen variants in a region where invasive disease by Salmonella Typhimurium is highly endemic. This needs to be taken into account when developing O-antigen-based vaccines, as it might impact the breadth of coverage in such regions. IMPORTANCE The bacterium Salmonella Typhimurium forms a devastating burden in sub-Saharan Africa by causing invasive bloodstream infections. Additionally, Salmonella Typhimurium presents high levels of antimicrobial resistance, jeopardizing treatment. No licensed vaccine is available, but candidates are in development, with lipopolysaccharides being the principal target of protective immunity. The vaccines under development are designed based on the O:5 antigen variant of bacterial lipopolysaccharides. Data on recent Salmonella Typhimurium clinical isolates from the Democratic Republic of the Congo (DRC), however, indicate increasing levels of isolates without this O:5 antigen. We studied this loss of O:5 antigen in the population at the genetic and chemical levels. We genome sequenced 354 isolates from the DRC and used advanced bioinformatics and chemical methods to characterize the lipopolysaccharide features within the bacterial population. Our results suggest evolutionary pressure toward O-antigen variants. This needs to be taken into account when developing vaccines, as it might impact vaccine coverage.
Keywords: Democratic Republic of the Congo; O-antigen; Salmonella Typhimurium; genomics; iNTS; surface structures; vaccines; whole-genome sequencing.
Conflict of interest statement
The authors declare a conflict of interest. GG, MMR and FM are employed by the GSK group of companies.
Figures


References
-
- Okoro CK, Kingsley RA, Connor TR, Harris SR, Parry CM, Al-Mashhadani MN, Kariuki S, Msefula CL, Gordon MA, de Pinna E, Wain J, Heyderman RS, Obaro S, Alonso PL, Mandomando I, MacLennan CA, Tapia MD, Levine MM, Tennant SM, Parkhill J, Dougan G. 2012. Intracontinental spread of human invasive Salmonella Typhimurium pathovariants in sub-Saharan Africa. Nat Genet 44:1215–1221. doi:10.1038/ng.2423. - DOI - PMC - PubMed
-
- Van Puyvelde S, Pickard D, Vandelannoote K, Heinz E, Barbe B, de Block T, Clare S, Coomber EL, Harcourt K, Sridhar S, Lees EA, Wheeler NE, Klemm EJ, Kuijpers L, Mbuyi Kalonji L, Phoba MF, Falay D, Ngbonda D, Lunguya O, Jacobs J, Dougan G, Deborggraeve S. 2019. An African Salmonella Typhimurium ST313 sublineage with extensive drug-resistance and signatures of host adaptation. Nat Commun 10:4280. doi:10.1038/s41467-019-11844-z. - DOI - PMC - PubMed
-
- Post AS, Diallo SN, Guiraud I, Lompo P, Tahita MC, Maltha J, Van Puyvelde S, Mattheus W, Ley B, Thriemer K, Rouamba E, Derra K, Deborggraeve S, Tinto H, Jacobs J. 2019. Supporting evidence for a human reservoir of invasive non-typhoidal Salmonella from household samples in Burkina Faso. PLoS Negl Trop Dis 13:e0007782. doi:10.1371/journal.pntd.0007782. - DOI - PMC - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Medical
Miscellaneous

