Dysregulation and prometastatic function of glycosyltransferase C1GALT1 modulated by cHP1BP3/ miR-1-3p axis in bladder cancer
- PMID: 35864552
- PMCID: PMC9306173
- DOI: 10.1186/s13046-022-02438-7
Dysregulation and prometastatic function of glycosyltransferase C1GALT1 modulated by cHP1BP3/ miR-1-3p axis in bladder cancer
Abstract
Background: Abnormal glycosylation in a variety of cancer types is involved in tumor progression and chemoresistance. Glycosyltransferase C1GALT1, the key enzyme in conversion of Tn antigen to T antigen, is involved in both physiological and pathological conditions. However, the mechanisms of C1GALT1 in enhancing oncogenic phenotypes and its regulatory effects via non-coding RNA are unclear.
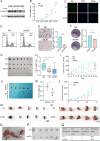
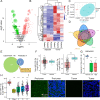
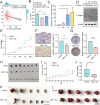
Methods: Abnormal expression of C1GALT1 and its products T antigen in human bladder cancer (BLCA) were evaluated with BLCA tissue, plasma samples and cell lines. Effects of C1GALT1 on migratory ability and proliferation were assessed in YTS-1 cells by transwell, CCK8 and colony formation assay in vitro and by mouse subcutaneous xenograft and trans-splenic metastasis models in vivo. Dysregulated circular RNAs (circRNAs) and microRNAs (miRNAs) were profiled in 3 pairs of bladder cancer tissues by RNA-seq. Effects of miR-1-3p and cHP1BP3 (circRNA derived from HP1BP3) on modulating C1GALT1 expression were investigated by target prediction program, correlation analysis and luciferase reporter assay. Functional roles of miR-1-3p and cHP1BP3 on migratory ability and proliferation in BLCA were also investigated by in vitro and in vivo experiments. Additionally, glycoproteomic analysis was employed to identify the target glycoproteins of C1GALT1.
Results: In this study, we demonstrated upregulation of C1GALT1 and its product T antigen in BLCA. C1GALT1 silencing suppressed migratory ability and proliferation of BLCA YTS-1 cells in vitro and in vivo. Subsets of circRNAs and miRNAs were dysregulated in BLCA tissues. miR-1-3p, which is reduced in BLCA tissues, inhibited transcription of C1GALT1 by binding directly to its 3'-untranslated region (3'-UTR). miR-1-3p overexpression resulted in decreased migratory ability and proliferation of YTS-1 cells. cHP1BP3 was upregulated in BLCA tissues, and served as an miR-1-3p "sponge". cHP1BP3 was shown to modulate migratory ability, proliferation, and colony formation of YTS-1 cells, and displayed tumor-suppressing activity in BLCA. Target glycoproteins of C1GALT1, including integrins and MUC16, were identified.
Conclusions: This study reveals the pro-metastatic and proliferative function of upregulated glycosyltransferase C1GLAT1, and provides preliminary data on mechanisms underlying dysregulation of C1GALT1 via miR-1-3p / cHP1BP3 axis in BLCA.
Keywords: Bladder cancer; C1GALT1; cHP1BP3; circRNA; miR-1-3p.
© 2022. The Author(s).
Conflict of interest statement
The authors declare no conflict of interest, or competing financial interest.
Figures





Similar articles
-
LncRNA MAFG-AS1 Promotes the Progression of Bladder Cancer by Targeting the miR-143-3p/COX-2 Axis.Pathobiology. 2020;87(6):345-355. doi: 10.1159/000509957. Epub 2020 Nov 25. Pathobiology. 2020. Retraction in: Pathobiology. 2025;92(3):187-188. doi: 10.1159/000544903. PMID: 33238264 Retracted.
-
Invasion-related circular RNA circFNDC3B inhibits bladder cancer progression through the miR-1178-3p/G3BP2/SRC/FAK axis.Mol Cancer. 2018 Nov 20;17(1):161. doi: 10.1186/s12943-018-0908-8. Mol Cancer. 2018. PMID: 30458784 Free PMC article.
-
Silencing circular RNA UVRAG inhibits bladder cancer growth and metastasis by targeting the microRNA-223/fibroblast growth factor receptor 2 axis.Cancer Sci. 2019 Jan;110(1):99-106. doi: 10.1111/cas.13857. Epub 2018 Dec 7. Cancer Sci. 2019. PMID: 30387298 Free PMC article.
-
The Tumor Suppressor Roles and Mechanisms of MiR-491 in Human Cancers.DNA Cell Biol. 2022 Sep;41(9):810-823. doi: 10.1089/dna.2022.0274. Epub 2022 Aug 1. DNA Cell Biol. 2022. PMID: 35914029 Review.
-
Role and mechanisms of noncoding RNAs in the regulation of metabolic reprogramming in bladder cancer (Review).Int J Mol Med. 2023 Sep;52(3):79. doi: 10.3892/ijmm.2023.5282. Epub 2023 Jul 21. Int J Mol Med. 2023. PMID: 37477143 Review.
Cited by
-
A serum panel of three microRNAs may serve as possible biomarkers for kidney renal clear cell carcinoma.Cancer Cell Int. 2024 Jan 8;24(1):18. doi: 10.1186/s12935-023-03187-z. Cancer Cell Int. 2024. PMID: 38191389 Free PMC article.
-
The important role of miR-1-3p in cancers.J Transl Med. 2023 Oct 31;21(1):769. doi: 10.1186/s12967-023-04649-8. J Transl Med. 2023. PMID: 37907984 Free PMC article. Review.
-
C1GALT1 induces the carcinogenesis of thyroid cancer through regulation by miR-141-3p and GLUT1.Heliyon. 2024 May 24;10(11):e31778. doi: 10.1016/j.heliyon.2024.e31778. eCollection 2024 Jun 15. Heliyon. 2024. PMID: 38845937 Free PMC article.
-
Small non-coding RNA profiling in patients with non-muscle invasive bladder cancer.BMC Cancer. 2025 Feb 21;25(1):319. doi: 10.1186/s12885-025-13672-5. BMC Cancer. 2025. PMID: 39984879 Free PMC article.
-
O-GlcNAcylation determines the function of the key O-GalNAc glycosyltransferase C1GalT1 in bladder cancer.Acta Biochim Biophys Sin (Shanghai). 2024 Aug 8;56(8):1108-1117. doi: 10.3724/abbs.2024129. Acta Biochim Biophys Sin (Shanghai). 2024. PMID: 39126245 Free PMC article.
References
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Medical
Molecular Biology Databases
Research Materials
Miscellaneous

