3D genome, on repeat: Higher-order folding principles of the heterochromatinized repetitive genome
- PMID: 35868274
- PMCID: PMC10225251
- DOI: 10.1016/j.cell.2022.06.052
3D genome, on repeat: Higher-order folding principles of the heterochromatinized repetitive genome
Abstract
Nearly half of the human genome is comprised of diverse repetitive sequences ranging from satellite repeats to retrotransposable elements. Such sequences are susceptible to stepwise expansions, duplications, inversions, and recombination events which can compromise genome function. In this review, we discuss the higher-order folding mechanisms of compartmentalization and loop extrusion and how they shape, and are shaped by, heterochromatin. Using primarily mammalian model systems, we contrast mechanisms governing H3K9me3-mediated heterochromatinization of the repetitive genome and highlight emerging links between repetitive elements and chromatin folding.
Copyright © 2022. Published by Elsevier Inc.
Conflict of interest statement
Declaration of interests The authors declare no competing interests.
Figures
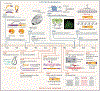




References
-
- Achour M, Jacq X, Rondé P, Alhosin M, Charlot C, Chataigneau T, Jeanblanc M, Macaluso M, Giordano A, Hughes AD, et al. (2008). The interaction of the SRA domain of ICBP90 with a novel domain of DNMT1 is involved in the regulation of VEGF gene expression. Oncogene 27, 2187–2197. 10.1038/sj.onc.1210855. - DOI - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

