GsMYB7 encoding a R2R3-type MYB transcription factor enhances the tolerance to aluminum stress in soybean (Glycine max L.)
- PMID: 35869448
- PMCID: PMC9306046
- DOI: 10.1186/s12864-022-08744-w
GsMYB7 encoding a R2R3-type MYB transcription factor enhances the tolerance to aluminum stress in soybean (Glycine max L.)
Abstract
Background: MYB transcription factor (TF) is one of the largest families of TFs in plants and play essential roles in plant growth and development, and is involved in responses to biological and abiotic stress. However, there are few reports on GsMYB7 gene in soybean under aluminum acid stress, and its regulatory mechanism remains unclear.
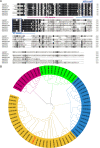
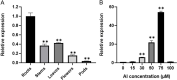
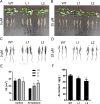
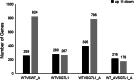
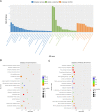
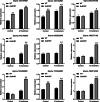
Results: The GsMYB7 protein is localized in the nucleus and has transcriptional activation ability. Quantitative real-time PCR (qRT-PCR) results showed that GsMYB7 held a constitutive expression pattern rich in roots. When AlCl3 concentration was 25 µM, the total root surface area (SA) of GsMYB7 transgenic lines were 34.97% higher than that of wild-type Huachun 6 (HC6). While the accumulation of Al3+ in root tip of transgenic plants after aluminum treatment was 17.39% lower than that of wild-type. RNA-sequencing analysis indicated that over 1181 genes were regulated by GsMYB7 and aluminum stress. Among all the regulated genes, the expression levels of glutathione peroxidase, protein kinase, cytochrome and other genes in the transgenic lines were significantly higher than those in wild type by acidic aluminum stress. The bioinformatics and qRT-PCR results showed that 9 candidate genes were induced under the treatments of acidic aluminum stress which were indirectly and/or directly regulated by GsMYB7. After AlCl3 treatments, the transcripts of these genes in GsMYB7 transgenic seedlings were significantly higher than those of wide-type HC6.
Conclusions: The results suggested that GsMYB7 may enhance soybean tolerance to acidic aluminum stress by regulating the downstream genes.
Keywords: Acidic aluminum; GsMYB7; R2R3-MYB; Soybean; Transcription factor.
© 2022. The Author(s).
Conflict of interest statement
The authors declare that they have no competing interests.
Figures



References
MeSH terms
Substances
Grants and funding
- CARS-04-PS09/the China Agricultural Research System
- 4100-C17106, 21301091702101/Ministry of Agriculture and Rural Affairs agricultural products quality and safety supervision special
- 2020B020220008/the Projects of Key Area Research and Development Program of Guangdong Province
- NZ2021012/the project of the Guangdong Provincial Laboratory of Lingnan Modern Agricultural Science and Technology
- 4100-31331/the Research Project of the State Key Laboratory of Agricultural and Biological Resources Protection and Utilization in Subtropics
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous

