Staggered intercalation of DNA duplexes with base-pair modulation by two distinct drug molecules induces asymmetric backbone twisting and structure polymorphism
- PMID: 35871296
- PMCID: PMC9410880
- DOI: 10.1093/nar/gkac629
Staggered intercalation of DNA duplexes with base-pair modulation by two distinct drug molecules induces asymmetric backbone twisting and structure polymorphism
Abstract
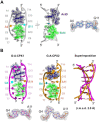
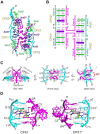
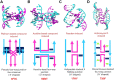
The use of multiple drugs simultaneously targeting DNA is a promising strategy in cancer therapy for potentially overcoming single drug resistance. In support of this concept, we report that a combination of actinomycin D (ActD) and echinomycin (Echi), can interact in novel ways with native and mismatched DNA sequences, distinct from the structural effects produced by either drug alone. Changes in the former with GpC and CpG steps separated by a A:G or G:A mismatch or in a native DNA with canonical G:C and C:G base pairs, result in significant asymmetric backbone twists through staggered intercalation and base pair modulations. A wobble or Watson-Crick base pair at the two drug-binding interfaces can result in a single-stranded 'chair-shaped' DNA duplex with a straight helical axis. However, a novel sugar-edged hydrogen bonding geometry in the G:A mismatch leads to a 'curved-shaped' duplex. Two non-canonical G:C Hoogsteen base pairings produce a sharply kinked duplex in different forms and a four-way junction-like superstructure, respectively. Therefore, single base pair modulations on the two drug-binding interfaces could significantly affect global DNA structure. These structures thus provide a rationale for atypical DNA recognition via multiple DNA intercalators and a structural basis for the drugs' potential synergetic use.
© The Author(s) 2022. Published by Oxford University Press on behalf of Nucleic Acids Research.
Figures




Similar articles
-
Atomic structures of excited state A-T Hoogsteen base pairs in duplex DNA by combining NMR relaxation dispersion, mutagenesis, and chemical shift calculations.J Biomol NMR. 2018 Apr;70(4):229-244. doi: 10.1007/s10858-018-0177-2. Epub 2018 Apr 19. J Biomol NMR. 2018. PMID: 29675775 Free PMC article.
-
Probing conformational transitions towards mutagenic Watson-Crick-like G·T mismatches using off-resonance sugar carbon R1ρ relaxation dispersion.J Biomol NMR. 2020 Sep;74(8-9):457-471. doi: 10.1007/s10858-020-00337-7. Epub 2020 Aug 12. J Biomol NMR. 2020. PMID: 32789613 Free PMC article.
-
NMR solution structure of a DNA-actinomycin D complex containing a non-hydrogen-bonding pair in the binding site.J Am Chem Soc. 2010 Dec 15;132(49):17588-98. doi: 10.1021/ja107575f. Epub 2010 Nov 22. J Am Chem Soc. 2010. PMID: 21090721
-
Watson-Crick versus Hoogsteen Base Pairs: Chemical Strategy to Encode and Express Genetic Information in Life.Acc Chem Res. 2021 May 4;54(9):2110-2120. doi: 10.1021/acs.accounts.0c00734. Epub 2021 Feb 16. Acc Chem Res. 2021. PMID: 33591181 Review.
-
Unusual DNA duplex and hairpin motifs.Nucleic Acids Res. 2003 May 15;31(10):2461-74. doi: 10.1093/nar/gkg367. Nucleic Acids Res. 2003. PMID: 12736295 Free PMC article. Review.
Cited by
-
Organometallic Pillarplexes That Bind DNA 4-Way Holliday Junctions and Forks.J Am Chem Soc. 2023 Jun 28;145(25):13570-13580. doi: 10.1021/jacs.3c00118. Epub 2023 Jun 15. J Am Chem Soc. 2023. PMID: 37318835 Free PMC article.
-
Dual dimeric interactions in the nucleic acid-binding protein Sac10b lead to multiple bridging of double-stranded DNA.Heliyon. 2024 May 23;10(11):e31630. doi: 10.1016/j.heliyon.2024.e31630. eCollection 2024 Jun 15. Heliyon. 2024. PMID: 38867953 Free PMC article.
-
Structural and Functional Insights into Targeting GCCG Sites in the EGFR Promoter by Two DNA Intercalators to Inhibit Breast Cancer Metastasis.J Med Chem. 2025 Mar 27;68(6):6601-6615. doi: 10.1021/acs.jmedchem.4c03203. Epub 2025 Mar 3. J Med Chem. 2025. PMID: 40032551 Free PMC article.
-
Synergistic binding of actinomycin D and echinomycin to DNA mismatch sites and their combined anti-tumour effects.Nucleic Acids Res. 2023 May 8;51(8):3540-3555. doi: 10.1093/nar/gkad156. Nucleic Acids Res. 2023. PMID: 36919604 Free PMC article.
-
The role of water in mediating DNA structures with epigenetic modifications, higher-order conformations and drug-DNA interactions.RSC Chem Biol. 2025 Mar 14;6(5):699-720. doi: 10.1039/d4cb00308j. eCollection 2025 May 8. RSC Chem Biol. 2025. PMID: 40171245 Free PMC article. Review.
References
-
- Granzhan A., Kotera N., Teulade-Fichou M.-P.. Finding needles in a basestack: recognition of mismatched base pairs in DNA by small molecules. Chem. Soc. Rev. 2014; 43:3630–3665. - PubMed
-
- Gibson D. Drug–DNA interactions and novel drug design. Pharmacogenomics J. 2002; 2:275–276. - PubMed
-
- Portugal J., Barceló F.. Noncovalent binding to DNA: still a target in developing anticancer agents. Curr. Med. Chem. 2016; 23:4108–4134. - PubMed
-
- Rescifina A., Zagni C., Varrica M.G., Pistarà V., Corsaro A.. Recent advances in small organic molecules as DNA intercalating agents: synthesis, activity, and modeling. Eur. J. Med. Chem. 2014; 74:95–115. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources

