The L1-ORF1p coiled coil enables formation of a tightly compacted nucleic acid-bound complex that is associated with retrotransposition
- PMID: 35871298
- PMCID: PMC9410894
- DOI: 10.1093/nar/gkac628
The L1-ORF1p coiled coil enables formation of a tightly compacted nucleic acid-bound complex that is associated with retrotransposition
Abstract
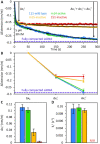
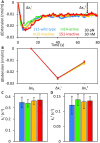
Long interspersed nuclear element 1 (L1) parasitized most vertebrates and constitutes ∼20% of the human genome. It encodes ORF1p and ORF2p which form an L1-ribonucleoprotein (RNP) with their encoding transcript that is copied into genomic DNA (retrotransposition). ORF1p binds single-stranded nucleic acid (ssNA) and exhibits NA chaperone activity. All vertebrate ORF1ps contain a coiled coil (CC) domain and we previously showed that a CC-retrotransposition null mutant prevented formation of stably bound ORF1p complexes on ssNA. Here, we compared CC variants using our recently improved method that measures ORF1p binding to ssDNA at different forces. Bound proteins decrease ssDNA contour length and at low force, retrotransposition-competent ORF1ps (111p and m14p) exhibit two shortening phases: the first is rapid, coincident with ORF1p binding; the second is slower, consistent with formation of tightly compacted complexes by NA-bound ORF1p. In contrast, two retrotransposition-null CC variants (151p and m15p) did not attain the second tightly compacted state. The C-terminal half of the ORF1p trimer (not the CC) contains the residues that mediate NA-binding. Our demonstrating that the CC governs the ability of NA-bound retrotransposition-competent trimers to form tightly compacted complexes reveals the biochemical phenotype of these coiled coil mutants.
© The Author(s) 2022. Published by Oxford University Press on behalf of Nucleic Acids Research.
Figures



Similar articles
-
Protein-nucleic acid interactions of LINE-1 ORF1p.Semin Cell Dev Biol. 2019 Feb;86:140-149. doi: 10.1016/j.semcdb.2018.03.019. Epub 2018 Mar 31. Semin Cell Dev Biol. 2019. PMID: 29596909 Free PMC article. Review.
-
L1-ORF1p nucleoprotein can rapidly assume distinct conformations and simultaneously bind more than one nucleic acid.Nucleic Acids Res. 2024 Dec 11;52(22):14013-14029. doi: 10.1093/nar/gkae1141. Nucleic Acids Res. 2024. PMID: 39565204 Free PMC article.
-
L1 retrotransposition requires rapid ORF1p oligomerization, a novel coiled coil-dependent property conserved despite extensive remodeling.Nucleic Acids Res. 2016 Jan 8;44(1):281-93. doi: 10.1093/nar/gkv1342. Epub 2015 Dec 15. Nucleic Acids Res. 2016. PMID: 26673717 Free PMC article.
-
Characterization of LINE-1 ribonucleoprotein particles.PLoS Genet. 2010 Oct 7;6(10):e1001150. doi: 10.1371/journal.pgen.1001150. PLoS Genet. 2010. PMID: 20949108 Free PMC article.
-
Nucleic acid chaperone properties of ORF1p from the non-LTR retrotransposon, LINE-1.RNA Biol. 2010 Nov-Dec;7(6):706-11. doi: 10.4161/rna.7.6.13766. Epub 2010 Nov 1. RNA Biol. 2010. PMID: 21045547 Free PMC article. Review.
Cited by
-
HIV-1 uncoating requires long double-stranded reverse transcription products.Sci Adv. 2024 Apr 26;10(17):eadn7033. doi: 10.1126/sciadv.adn7033. Epub 2024 Apr 24. Sci Adv. 2024. PMID: 38657061 Free PMC article.
-
Diverse single-stranded nucleic acid binding proteins enable both stable protection and rapid exchange required for biological function.QRB Discov. 2025 Jan 14;6:e1. doi: 10.1017/qrd.2024.21. eCollection 2025. QRB Discov. 2025. PMID: 39944884 Free PMC article.
-
LINE-1 ribonucleoprotein condensates bind DNA to enable nuclear entry during mitosis.Sci Adv. 2025 May 2;11(18):eadt9318. doi: 10.1126/sciadv.adt9318. Epub 2025 May 2. Sci Adv. 2025. PMID: 40315332 Free PMC article.
-
Dynamic structure of T4 gene 32 protein filaments facilitates rapid noncooperative protein dissociation.Nucleic Acids Res. 2023 Sep 8;51(16):8587-8605. doi: 10.1093/nar/gkad595. Nucleic Acids Res. 2023. PMID: 37449435 Free PMC article.
-
DNA damage alters binding conformations of E. coli single-stranded DNA-binding protein.Biophys J. 2023 Oct 3;122(19):3950-3958. doi: 10.1016/j.bpj.2023.08.018. Epub 2023 Aug 24. Biophys J. 2023. PMID: 37632138 Free PMC article.
References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

