Targeting ACLY efficiently inhibits SARS-CoV-2 replication
- PMID: 35874959
- PMCID: PMC9305265
- DOI: 10.7150/ijbs.72709
Targeting ACLY efficiently inhibits SARS-CoV-2 replication
Abstract
The Coronavirus Disease 2019 (COVID-19) pandemic caused by severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) is the biggest public health challenge the world has witnessed in the past decades. SARS-CoV-2 undergoes constant mutations and new variants of concerns (VOCs) with altered transmissibility, virulence, and/or susceptibility to vaccines and therapeutics continue to emerge. Detailed analysis of host factors involved in virus replication may help to identify novel treatment targets. In this study, we dissected the metabolome derived from COVID-19 patients to identify key host factors that are required for efficient SARS-CoV-2 replication. Through a series of metabolomic analyses, in vitro, and in vivo investigations, we identified ATP citrate lyase (ACLY) as a novel host factor required for efficient replication of SARS-CoV-2 wild-type and variants, including Omicron. ACLY should be further explored as a novel intervention target for COVID-19.
Keywords: ACLY; COVID-19; Delta; Omicron; SARS-CoV-2; metabolomics.
© The author(s).
Conflict of interest statement
Competing Interests: J.F.-W.C. has received travel grants from Pfizer Corporation Hong Kong and Astellas Pharma Hong Kong Corporation Limited and was an invited speaker for Gilead Sciences Hong Kong Limited and Luminex Corporation. The other authors declare no competing interests.
Figures
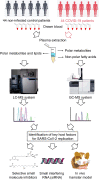






References
-
- WHO. Weekly operational update on COVID-19 - 1 March 2022. https://www.who.int/publications/m/item/weekly-operational-update-on-cov....
-
- Shuai H, Chan JF, Hu B, Chai Y, Yuen TT, Yin F. et al. Attenuated replication and pathogenicity of SARS-CoV-2 B.1.1.529 Omicron. Nature. 2022;603:693–9. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Medical
Miscellaneous

