Computational Characterizing Necroptosis Reveals Implications for Immune Infiltration and Immunotherapy of Hepatocellular Carcinoma
- PMID: 35875102
- PMCID: PMC9301124
- DOI: 10.3389/fonc.2022.933210
Computational Characterizing Necroptosis Reveals Implications for Immune Infiltration and Immunotherapy of Hepatocellular Carcinoma
Abstract
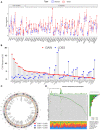
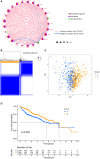
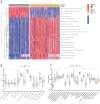
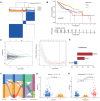
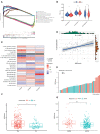
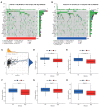
Necroptosis is a programmed form of necrotic cell death in regulating cancer ontogenesis, progression, and tumor microenvironment (TME) and could drive tumor-infiltrating cells to release pro-inflammatory cytokines, incurring strong immune responses. Nowadays, there are few identified biomarkers applied in clinical immunotherapy, and it is increasingly recognized that high levels of tumor necroptosis could enhance the response to immunotherapy. However, comprehensive characterization of necroptosis associated with TME and immunotherapy in Hepatocellular carcinoma (HCC) remains unexplored. Here, we computationally characterized necroptosis landscape in HCC samples from TCGA and ICGA cohorts and stratified them into two necroptosis clusters (A or B) with significantly different characteristics in clinical prognosis, immune cell function, and TME-landscapes. Additionally, to further evaluate the necroptosis levels of each sample, we established a novel necroptosis-related gene score (NRGscore). We further investigated the TME, tumor mutational burden (TMB), clinical response to immunotherapy, and chemotherapeutic drug sensitivity of HCC subgroups stratified by the necroptosis landscapes. The NRGscore is robust and highly predictive of HCC clinical outcomes. Further analysis indicated that the high NRGscore group resembles the immune-inflamed phenotype while the low score group is analogous to the immune-exclusion or metabolism phenotype. Additionally, the high NRGscore group is more sensitive to immune checkpoint blockade-based immunotherapy, which was further validated using an external HCC cohort, metastatic melanoma cohort, and advanced urothelial cancer cohort. Besides, the NRGscore was demonstrated as a potential biomarker for chemotherapy, wherein the high NRGscore patients with more tumor stem cell composition could be more sensitive to Cisplatin, Doxorubicin, Paclitaxel-based chemotherapy, and Sorafenib therapy. Collectively, a comprehensive characterization of the necroptosis in HCC suggested its implications for predicting immune infiltration and response to immunotherapy of HCC, providing promising strategies for treatment.
Keywords: necroptosis; chemotherapy; hepatocellular carcinoma; immunotherapy; tumor microenvironment; tumor-infiltrating cells.
Copyright © 2022 Zhu, Han, Zhao, Zhu, Ma, Xu, Bai, Tang, Xu and Liu.
Conflict of interest statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Figures

Similar articles
-
A cellular senescence-related classifier based on a tumorigenesis- and immune infiltration-guided strategy can predict prognosis, immunotherapy response, and candidate drugs in hepatocellular carcinoma.Front Immunol. 2022 Nov 15;13:974377. doi: 10.3389/fimmu.2022.974377. eCollection 2022. Front Immunol. 2022. PMID: 36458010 Free PMC article.
-
A Novel Prognostic Signature Associated with Immunotherapeutic Response for Hepatocellular Carcinoma.Front Surg. 2022 Jul 5;9:905897. doi: 10.3389/fsurg.2022.905897. eCollection 2022. Front Surg. 2022. PMID: 35865037 Free PMC article.
-
Integrated single cell and bulk sequencing analysis identifies tumor reactive CXCR6+ CD8 T cells as a predictor of immune infiltration and immunotherapy outcomes in hepatocellular carcinoma.Front Oncol. 2023 Aug 1;13:1099385. doi: 10.3389/fonc.2023.1099385. eCollection 2023. Front Oncol. 2023. PMID: 37593098 Free PMC article.
-
A novel necroptosis-related gene index for predicting prognosis and a cold tumor immune microenvironment in stomach adenocarcinoma.Front Immunol. 2022 Oct 27;13:968165. doi: 10.3389/fimmu.2022.968165. eCollection 2022. Front Immunol. 2022. PMID: 36389725 Free PMC article. Review.
-
Strategies to Improve the Antitumor Effect of Immunotherapy for Hepatocellular Carcinoma.Front Immunol. 2021 Nov 26;12:783236. doi: 10.3389/fimmu.2021.783236. eCollection 2021. Front Immunol. 2021. PMID: 34899747 Free PMC article. Review.
Cited by
-
Modulating the Cell Death Immune Response for Electroporation Treatments.Bioelectricity. 2024 Dec 13;6(4):263-271. doi: 10.1089/bioe.2024.0019. eCollection 2024 Dec. Bioelectricity. 2024. PMID: 39712216
References
-
- Amin MB, Greene FL, Edge SB, Compton CC, Gershenwald JE, Brookland RK, et al. . The Eighth Edition AJCC Cancer Staging Manual: Continuing to Build a Bridge From a Population-Based to a More "Personalized" Approach to Cancer Staging. CA: Cancer J Clin (2017) 67:93–9. doi: 10.3322/caac.21388 - DOI - PubMed
LinkOut - more resources
Full Text Sources

