Constraint-Based Reconstruction and Analyses of Metabolic Models: Open-Source Python Tools and Applications to Cancer
- PMID: 35875150
- PMCID: PMC9303011
- DOI: 10.3389/fonc.2022.914594
Constraint-Based Reconstruction and Analyses of Metabolic Models: Open-Source Python Tools and Applications to Cancer
Abstract
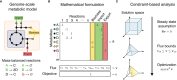
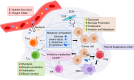
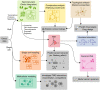
The influence of metabolism on signaling, epigenetic markers, and transcription is highly complex yet important for understanding cancer physiology. Despite the development of high-resolution multi-omics technologies, it is difficult to infer metabolic activity from these indirect measurements. Fortunately, genome-scale metabolic models and constraint-based modeling provide a systems biology framework to investigate the metabolic states and define the genotype-phenotype associations by integrations of multi-omics data. Constraint-Based Reconstruction and Analysis (COBRA) methods are used to build and simulate metabolic networks using mathematical representations of biochemical reactions, gene-protein reaction associations, and physiological and biochemical constraints. These methods have led to advancements in metabolic reconstruction, network analysis, perturbation studies as well as prediction of metabolic state. Most computational tools for performing these analyses are written for MATLAB, a proprietary software. In order to increase accessibility and handle more complex datasets and models, community efforts have started to develop similar open-source tools in Python. To date there is a comprehensive set of tools in Python to perform various flux analyses and visualizations; however, there are still missing algorithms in some key areas. This review summarizes the availability of Python software for several components of COBRA methods and their applications in cancer metabolism. These tools are evolving rapidly and should offer a readily accessible, versatile way to model the intricacies of cancer metabolism for identifying cancer-specific metabolic features that constitute potential drug targets.
Keywords: cancer; constraint-based modeling; genome-scale metabolic models; metabolism; omics; python; single-cell analysis; systems biology.
Copyright © 2022 Ng, Lee, Baloni, Diener, Heath and Su.
Conflict of interest statement
JH is a board member of PACT Pharma and Isoplexis and receives support from Gilead, Regeneron and Merck. The remaining authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Figures

Similar articles
-
A pipeline for the reconstruction and evaluation of context-specific human metabolic models at a large-scale.PLoS Comput Biol. 2022 Jun 24;18(6):e1009294. doi: 10.1371/journal.pcbi.1009294. eCollection 2022 Jun. PLoS Comput Biol. 2022. PMID: 35749559 Free PMC article.
-
COBRApy: COnstraints-Based Reconstruction and Analysis for Python.BMC Syst Biol. 2013 Aug 8;7:74. doi: 10.1186/1752-0509-7-74. BMC Syst Biol. 2013. PMID: 23927696 Free PMC article.
-
A simplified metabolic network reconstruction to promote understanding and development of flux balance analysis tools.Comput Biol Med. 2019 Feb;105:64-71. doi: 10.1016/j.compbiomed.2018.12.010. Epub 2018 Dec 17. Comput Biol Med. 2019. PMID: 30584952
-
Computational Modeling of Human Metabolism and Its Application to Systems Biomedicine.Methods Mol Biol. 2016;1386:253-81. doi: 10.1007/978-1-4939-3283-2_12. Methods Mol Biol. 2016. PMID: 26677187 Review.
-
Machine learning for the advancement of genome-scale metabolic modeling.Biotechnol Adv. 2024 Sep;74:108400. doi: 10.1016/j.biotechadv.2024.108400. Epub 2024 Jun 27. Biotechnol Adv. 2024. PMID: 38944218 Review.
Cited by
-
Applying Proteomics and Computational Approaches to Identify Novel Targets in Blast-Associated Post-Traumatic Epilepsy.Int J Mol Sci. 2024 Mar 1;25(5):2880. doi: 10.3390/ijms25052880. Int J Mol Sci. 2024. PMID: 38474127 Free PMC article.
-
Integrating multi-omics to unravel host-microbiome interactions in inflammatory bowel disease.Cell Rep Med. 2024 Sep 17;5(9):101738. doi: 10.1016/j.xcrm.2024.101738. Cell Rep Med. 2024. PMID: 39293401 Free PMC article. Review.
-
Adding metabolic tasks to human GEM models to improve the study of gene targets and their associated toxicities.Sci Rep. 2024 Jul 27;14(1):17265. doi: 10.1038/s41598-024-68073-8. Sci Rep. 2024. PMID: 39068208 Free PMC article.
-
Metabolic model predictions enable targeted microbiome manipulation through precision prebiotics.bioRxiv [Preprint]. 2023 Feb 18:2023.02.17.528811. doi: 10.1101/2023.02.17.528811. bioRxiv. 2023. Update in: Microbiol Spectr. 2024 Feb 6;12(2):e0114423. doi: 10.1128/spectrum.01144-23. PMID: 36824941 Free PMC article. Updated. Preprint.
-
Kinetic Trajectories of Glucose Uptake in Single Cancer Cells Reveal a Drug-Induced Cell-State Change Within Hours of Drug Treatment.J Phys Chem B. 2024 Aug 22;128(33):7978-7986. doi: 10.1021/acs.jpcb.4c03663. Epub 2024 Aug 8. J Phys Chem B. 2024. PMID: 39115241 Free PMC article.
References
Publication types
LinkOut - more resources
Full Text Sources

