Rearrangement with the nkd2 promoter contributed to allelic diversity of the r1 gene in maize (Zea mays)
- PMID: 35876146
- PMCID: PMC9546038
- DOI: 10.1111/tpj.15918
Rearrangement with the nkd2 promoter contributed to allelic diversity of the r1 gene in maize (Zea mays)
Abstract
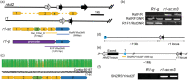
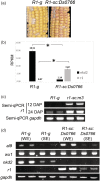
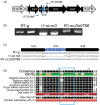
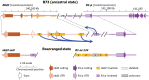
The maize red1 (r1) locus regulates anthocyanin accumulation and is a classic model for allelic diversity; changes in regulatory regions are responsible for most of the variation in gene expression patterns. Here, an intrachromosomal rearrangement between the distal upstream region of r1 and the region of naked endosperm 2 (nkd2) upstream to the third exon generated a nkd2 null allele lacking the first three exons, and the R1-st (stippled) allele with a novel r1 5' promoter region homologous to 5' regions from nkd2-B73. R1-sc:124 (an R1-st derivative) shows increased and earlier expression than a standard R1-g allele, as well as ectopic expression in the starchy endosperm compartment. Laser capture microdissection and RNA sequencing indicated that ectopic R1-sc:124 expression impacted expression of genes associated with RNA modification. The expression of R1-sc:124 resembled nkd2-W22 expression, suggesting that nkd2 regulatory sequences may influence the expression of R1-sc:124. The r1-sc:m3 allele is derived from R1-sc:124 by an insertion of a Ds6 transposon in intron 4. This insertion blocks anthocyanin regulation by causing mis-splicing that eliminates exon 5 from the mRNA. This allele serves as an important launch site for Ac/Ds mutagenesis studies, and two Ds6 insertions believed to be associated with nkd2 mutant alleles were actually located in the r1 5' region. Among annotated genomes of teosinte and maize varieties, the nkd2 and r1 loci showed conserved overall gene structures, similar to the B73 reference genome, suggesting that the nkd2-r1 rearrangement may be a recent event.
Keywords: Zea mays; aleurone; anthocyanin; inversion; kernel.
© 2022 The Authors. The Plant Journal published by Society for Experimental Biology and John Wiley & Sons Ltd.
Conflict of interest statement
The authors declare no conflicts of interest with this research.
Figures




Similar articles
-
NKD Transcription Factors Are Central Regulators of Maize Endosperm Development.Plant Cell. 2016 Dec;28(12):2916-2936. doi: 10.1105/tpc.16.00609. Epub 2016 Nov 28. Plant Cell. 2016. PMID: 27895224 Free PMC article.
-
A newly characterized allele of ZmR1 increases anthocyanin content in whole maize plant and the regulation mechanism of different ZmR1 alleles.Theor Appl Genet. 2022 Sep;135(9):3039-3055. doi: 10.1007/s00122-022-04166-0. Epub 2022 Jul 5. Theor Appl Genet. 2022. PMID: 35788748
-
The naked endosperm genes encode duplicate INDETERMINATE domain transcription factors required for maize endosperm cell patterning and differentiation.Plant Physiol. 2015 Feb;167(2):443-56. doi: 10.1104/pp.114.251413. Epub 2014 Dec 31. Plant Physiol. 2015. PMID: 25552497 Free PMC article.
-
Gene conversion within regulatory sequences generates maize r alleles with altered gene expression.Genetics. 2001 Dec;159(4):1727-40. doi: 10.1093/genetics/159.4.1727. Genetics. 2001. PMID: 11779810 Free PMC article.
-
Maize endosperm development.J Integr Plant Biol. 2021 Apr;63(4):613-627. doi: 10.1111/jipb.13069. Epub 2021 Mar 8. J Integr Plant Biol. 2021. PMID: 33448626 Review.
References
-
- Ahern, K.R. , Deewatthanawong, P. , Schares, J. , Muszynski, M. , Weeks, R. , Vollbrecht, E. et al. (2009) Regional mutagenesis using dissociation in maize. Methods, 49, 248–254. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

