Biosynthesis of uridine diphosphate N-Acetylglucosamine: An underexploited pathway in the search for novel antibiotics?
- PMID: 35880704
- PMCID: PMC10087520
- DOI: 10.1002/iub.2664
Biosynthesis of uridine diphosphate N-Acetylglucosamine: An underexploited pathway in the search for novel antibiotics?
Abstract
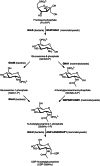
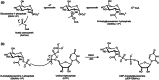
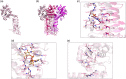
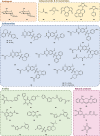
Although the prevalence of antibiotic resistance is increasing at an alarming rate, there are a dwindling number of effective antibiotics available. Thus, the development of novel antibacterial agents should be of utmost importance. Peptidoglycan biosynthesis has been and is still an attractive source for antibiotic targets; however, there are several components that remain underexploited. In this review, we examine the enzymes involved in the biosynthesis of one such component, UDP-N-acetylglucosamine, an essential building block and precursor of bacterial peptidoglycan. Furthermore, given the presence of a similar biosynthesis pathway in eukaryotes, we discuss the current knowledge on the differences and similarities between the bacterial and eukaryotic enzymes. Finally, this review also summarises the recent advances made in the development of inhibitors targeting the bacterial enzymes.
Keywords: UDP-N-acetylglucosamine biosynthesis; antibiotic resistance; bifunctional GlmU; glucosamine-6-phosphate synthase; peptidoglycan biosynthesis; phosphoglucosamine mutase.
© 2022 The Authors. IUBMB Life published by Wiley Periodicals LLC on behalf of International Union of Biochemistry and Molecular Biology.
Figures






Similar articles
-
Copurification of glucosamine-1-phosphate acetyltransferase and N-acetylglucosamine-1-phosphate uridyltransferase activities of Escherichia coli: characterization of the glmU gene product as a bifunctional enzyme catalyzing two subsequent steps in the pathway for UDP-N-acetylglucosamine synthesis.J Bacteriol. 1994 Sep;176(18):5788-95. doi: 10.1128/jb.176.18.5788-5795.1994. J Bacteriol. 1994. PMID: 8083170 Free PMC article.
-
Biosynthesis of wall polymers in Bacillus subtilis.J Bacteriol. 1977 Jun;130(3):1055-63. doi: 10.1128/jb.130.3.1055-1063.1977. J Bacteriol. 1977. PMID: 405370 Free PMC article.
-
Acetyltransfer precedes uridylyltransfer in the formation of UDP-N-acetylglucosamine in separable active sites of the bifunctional GlmU protein of Escherichia coli.Biochemistry. 1996 Jan 16;35(2):579-85. doi: 10.1021/bi952275a. Biochemistry. 1996. PMID: 8555230
-
Insights into the central role of N-acetyl-glucosamine-1-phosphate uridyltransferase (GlmU) in peptidoglycan metabolism and its potential as a therapeutic target.Biochem J. 2023 Jun 18;480(15):1147-1164. doi: 10.1042/BCJ20230173. Biochem J. 2023. PMID: 37498748 Review.
-
Cytoplasmic steps of peptidoglycan biosynthesis.FEMS Microbiol Rev. 2008 Mar;32(2):168-207. doi: 10.1111/j.1574-6976.2008.00104.x. Epub 2008 Feb 11. FEMS Microbiol Rev. 2008. PMID: 18266853 Review.
Cited by
-
A special issue of IUBMB Life celebrating the 50th anniversary of FAOBMB (1972-2022).IUBMB Life. 2022 Dec;74(12):1124-1125. doi: 10.1002/iub.2691. Epub 2022 Nov 22. IUBMB Life. 2022. PMID: 36345606 Free PMC article. No abstract available.
-
Isocyanides inhibit bacterial pathogens by covalent targeting of essential metabolic enzymes.Chem Sci. 2024 Jun 26;15(30):11946-11955. doi: 10.1039/d4sc01940g. eCollection 2024 Jul 31. Chem Sci. 2024. PMID: 39092115 Free PMC article.
-
Deconvolution of multichannel LC-MS/MS chromatograms of glucosamine-phosphates: Evidence of a GlmS regulatory difference between Staphylococcus aureus and Enterococcus faecium.Talanta Open. 2023 Dec;8:100241. doi: 10.1016/j.talo.2023.100241. Epub 2023 Jun 12. Talanta Open. 2023. PMID: 38187186 Free PMC article.
-
The Hexosamine Biosynthesis Pathway: Regulation and Function.Genes (Basel). 2023 Apr 18;14(4):933. doi: 10.3390/genes14040933. Genes (Basel). 2023. PMID: 37107691 Free PMC article. Review.
-
Gfa1 (glutamine fructose-6-phosphate aminotransferase) is essential for Aspergillus fumigatus growth and virulence.BMC Biol. 2025 Mar 13;23(1):80. doi: 10.1186/s12915-025-02184-0. BMC Biol. 2025. PMID: 40082985 Free PMC article.
References
-
- Chopra I, Schofield C, Everett M, O'Neill A, Miller K, et al. Treatment of health‐care‐associated infections caused by gram‐negative bacteria: A consensus statement. Lancet Infect Dis. 2008;8:133–139. - PubMed
-
- Centers for Disease Control and Prevention . Antibiotic resistance threats in the United States, 2019. Georgia: Atlanta, 2019.
-
- O'Neill, J. (2016) The review on antimicrobial resistance. Tackling drug‐resistant infections globally: Final report and recommendations. London, UK.
-
- Tacconelli E, Carrara E, Savoldi A, et al. Discovery, research, and development of new antibiotics: The WHO priority list of antibiotic‐resistant bacteria and tuberculosis. Lancet Infect Dis. 2018;18:318–327. - PubMed
-
- Rice LB. Federal funding for the study of antimicrobial resistance in nosocomial pathogens: No ESKAPE. J Infect Dis. 2008;197:1079–1081. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Medical
Molecular Biology Databases

