Revealing the Dynamics of Neural Information Processing with Multivariate Information Decomposition
- PMID: 35885153
- PMCID: PMC9319160
- DOI: 10.3390/e24070930
Revealing the Dynamics of Neural Information Processing with Multivariate Information Decomposition
Abstract
The varied cognitive abilities and rich adaptive behaviors enabled by the animal nervous system are often described in terms of information processing. This framing raises the issue of how biological neural circuits actually process information, and some of the most fundamental outstanding questions in neuroscience center on understanding the mechanisms of neural information processing. Classical information theory has long been understood to be a natural framework within which information processing can be understood, and recent advances in the field of multivariate information theory offer new insights into the structure of computation in complex systems. In this review, we provide an introduction to the conceptual and practical issues associated with using multivariate information theory to analyze information processing in neural circuits, as well as discussing recent empirical work in this vein. Specifically, we provide an accessible introduction to the partial information decomposition (PID) framework. PID reveals redundant, unique, and synergistic modes by which neurons integrate information from multiple sources. We focus particularly on the synergistic mode, which quantifies the "higher-order" information carried in the patterns of multiple inputs and is not reducible to input from any single source. Recent work in a variety of model systems has revealed that synergistic dynamics are ubiquitous in neural circuitry and show reliable structure-function relationships, emerging disproportionately in neuronal rich clubs, downstream of recurrent connectivity, and in the convergence of correlated activity. We draw on the existing literature on higher-order information dynamics in neuronal networks to illustrate the insights that have been gained by taking an information decomposition perspective on neural activity. Finally, we briefly discuss future promising directions for information decomposition approaches to neuroscience, such as work on behaving animals, multi-target generalizations of PID, and time-resolved local analyses.
Keywords: computation; cortical circuits; entropy; higher-order interactions; information theory; neural recording; neuroscience.
Conflict of interest statement
The authors declare no conflict of interest.
Figures
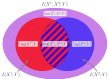



References
-
- Williams P.L., Beer R.D. Nonnegative Decomposition of Multivariate Information. arXiv. 20101004.2515
-
- Sherrill S.P., Timme N.M., Beggs J.M., Newman E.L. Partial information decomposition reveals that synergistic neural integration is greater downstream of recurrent information flow in organotypic cortical cultures. PLoS Comput. Biol. 2021;17:e1009196. doi: 10.1371/journal.pcbi.1009196. - DOI - PMC - PubMed
Publication types
Grants and funding
LinkOut - more resources
Full Text Sources

