Current Status of Regulatory Non-Coding RNAs Research in the Tritryp
- PMID: 35893237
- PMCID: PMC9326685
- DOI: 10.3390/ncrna8040054
Current Status of Regulatory Non-Coding RNAs Research in the Tritryp
Abstract
Trypanosomatids are protozoan parasites that cause devastating vector-borne human diseases. Gene expression regulation of these organisms depends on post-transcriptional control in responding to diverse environments while going through multiple developmental stages of their complex life cycles. In this scenario, non-coding RNAs (ncRNAs) are excellent candidates for a very efficient, quick, and economic strategy to regulate gene expression. The advent of high throughput RNA sequencing technologies show the presence and deregulation of small RNA fragments derived from canonical ncRNAs. This review seeks to depict the ncRNA landscape in trypanosomatids, focusing on the small RNA fragments derived from functional RNA molecules observed in RNA sequencing studies. Small RNA fragments derived from canonical ncRNAs (tsRNAs, snsRNAs, sdRNAs, and sdrRNAs) were identified in trypanosomatids. Some of these RNAs display changes in their levels associated with different environments and developmental stages, demanding further studies to determine their functional characterization and potential roles. Nevertheless, a comprehensive and detailed ncRNA annotation for most trypanosomatid genomes is still needed, allowing better and more extensive comparative and functional studies.
Keywords: Trypanosoma; leishmania; ncRNA; non-coding RNA; siRNA; snRNA; snoRNA; tRNA; tritryp; trypanosomatids.
Conflict of interest statement
The authors declare no conflict of interest.
Figures
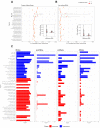

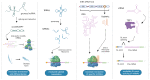
Similar articles
-
A comparative genome-wide study of ncRNAs in trypanosomatids.BMC Genomics. 2010 Nov 4;11:615. doi: 10.1186/1471-2164-11-615. BMC Genomics. 2010. PMID: 21050447 Free PMC article.
-
Genome-wide computational identification of functional RNA elements in Trypanosoma brucei.BMC Genomics. 2009 Aug 4;10:355. doi: 10.1186/1471-2164-10-355. BMC Genomics. 2009. PMID: 19653906 Free PMC article.
-
Epitranscriptome machinery in Trypanosomatids: New players on the table?Mol Microbiol. 2021 May;115(5):942-958. doi: 10.1111/mmi.14688. Epub 2021 Feb 10. Mol Microbiol. 2021. PMID: 33513291
-
tRNA-Derived Short Non-coding RNA as Interacting Partners of Argonaute Proteins.Gene Regul Syst Bio. 2015 Sep 10;9:27-33. doi: 10.4137/GRSB.S29411. eCollection 2015. Gene Regul Syst Bio. 2015. PMID: 26401098 Free PMC article. Review.
-
Global survey of miRNAs and tRNA-derived small RNAs from the human parasitic protist Trichomonas vaginalis.Parasit Vectors. 2021 Jan 29;14(1):87. doi: 10.1186/s13071-020-04570-9. Parasit Vectors. 2021. PMID: 33514387 Free PMC article.
Cited by
-
Nanopore-Based Direct RNA Sequencing of the Trypanosoma brucei Transcriptome Identifies Novel lncRNAs.Genes (Basel). 2023 Feb 28;14(3):610. doi: 10.3390/genes14030610. Genes (Basel). 2023. PMID: 36980882 Free PMC article.
-
The Role of Hydrogen Sulfide (H2S) in Epigenetic Regulation of Neurodegenerative Diseases: A Systematic Review.Int J Mol Sci. 2023 Aug 8;24(16):12555. doi: 10.3390/ijms241612555. Int J Mol Sci. 2023. PMID: 37628735 Free PMC article.
-
Comparative and systems analyses of Leishmania spp. non-coding RNAs through developmental stages.PLoS Negl Trop Dis. 2025 May 28;19(5):e0013108. doi: 10.1371/journal.pntd.0013108. eCollection 2025 May. PLoS Negl Trop Dis. 2025. PMID: 40435329 Free PMC article.
-
Life in plastic, it's fantastic! How Leishmania exploit genome instability to shape gene expression.Front Cell Infect Microbiol. 2023 Jan 26;13:1102462. doi: 10.3389/fcimb.2023.1102462. eCollection 2023. Front Cell Infect Microbiol. 2023. PMID: 36779182 Free PMC article. Review.
-
Systematic computational hunting for small RNAs derived from ncRNAs during dengue virus infection in endothelial HMEC-1 cells.Front Bioinform. 2024 Jan 31;4:1293412. doi: 10.3389/fbinf.2024.1293412. eCollection 2024. Front Bioinform. 2024. PMID: 38357577 Free PMC article.
References
Publication types
Grants and funding
LinkOut - more resources
Full Text Sources

