Model-based stationarity filtering of long-term memory data applied to resting-state blood-oxygen-level-dependent signal
- PMID: 35895686
- PMCID: PMC9328502
- DOI: 10.1371/journal.pone.0268752
Model-based stationarity filtering of long-term memory data applied to resting-state blood-oxygen-level-dependent signal
Abstract
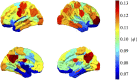
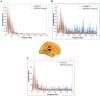
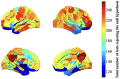
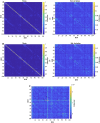
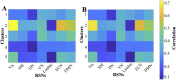
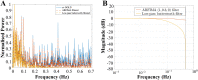
Resting-state blood-oxygen-level-dependent (BOLD) signal acquired through functional magnetic resonance imaging is a proxy of neural activity and a key mechanism for assessing neurological conditions. Therefore, practical tools to filter out artefacts that can compromise the assessment are required. On the one hand, a variety of tailored methods to preprocess the data to deal with identified sources of noise (e.g., head motion, heart beating, and breathing, just to mention a few) are in place. But, on the other hand, there might be unknown sources of unstructured noise present in the data. Therefore, to mitigate the effects of such unstructured noises, we propose a model-based filter that explores the statistical properties of the underlying signal (i.e., long-term memory). Specifically, we consider autoregressive fractional integrative process filters. Remarkably, we provide evidence that such processes can model the signals at different regions of interest to attain stationarity. Furthermore, we use a principled analysis where a ground-truth signal with statistical properties similar to the BOLD signal under the injection of noise is retrieved using the proposed filters. Next, we considered preprocessed (i.e., the identified sources of noise removed) resting-state BOLD data of 98 subjects from the Human Connectome Project. Our results demonstrate that the proposed filters decrease the power in the higher frequencies. However, unlike the low-pass filters, the proposed filters do not remove all high-frequency information, instead they preserve process-related higher frequency information. Additionally, we considered four different metrics (power spectrum, functional connectivity using the Pearson's correlation, coherence, and eigenbrains) to infer the impact of such filter. We provided evidence that whereas the first three keep most of the features of interest from a neuroscience perspective unchanged, the latter exhibits some variations that could be due to the sporadic activity filtered out.
Conflict of interest statement
The authors have declared that no competing interests exist.
Figures






References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

